Filter
Associated Lab
- Aguilera Castrejon Lab (19) Apply Aguilera Castrejon Lab filter
- Ahrens Lab (73) Apply Ahrens Lab filter
- Aso Lab (42) Apply Aso Lab filter
- Baker Lab (38) Apply Baker Lab filter
- Betzig Lab (116) Apply Betzig Lab filter
- Beyene Lab (15) Apply Beyene Lab filter
- Bock Lab (17) Apply Bock Lab filter
- Branson Lab (55) Apply Branson Lab filter
- Card Lab (43) Apply Card Lab filter
- Cardona Lab (64) Apply Cardona Lab filter
- Chklovskii Lab (13) Apply Chklovskii Lab filter
- Clapham Lab (16) Apply Clapham Lab filter
- Cui Lab (19) Apply Cui Lab filter
- Darshan Lab (12) Apply Darshan Lab filter
- Dennis Lab (3) Apply Dennis Lab filter
- Dickson Lab (46) Apply Dickson Lab filter
- Druckmann Lab (25) Apply Druckmann Lab filter
- Dudman Lab (56) Apply Dudman Lab filter
- Eddy/Rivas Lab (30) Apply Eddy/Rivas Lab filter
- Egnor Lab (11) Apply Egnor Lab filter
- Espinosa Medina Lab (23) Apply Espinosa Medina Lab filter
- Feliciano Lab (12) Apply Feliciano Lab filter
- Fetter Lab (41) Apply Fetter Lab filter
- FIB-SEM Technology (1) Apply FIB-SEM Technology filter
- Fitzgerald Lab (30) Apply Fitzgerald Lab filter
- Freeman Lab (15) Apply Freeman Lab filter
- Funke Lab (46) Apply Funke Lab filter
- Gonen Lab (91) Apply Gonen Lab filter
- Grigorieff Lab (62) Apply Grigorieff Lab filter
- Harris Lab (65) Apply Harris Lab filter
- Heberlein Lab (94) Apply Heberlein Lab filter
- Hermundstad Lab (30) Apply Hermundstad Lab filter
- Hess Lab (80) Apply Hess Lab filter
- Ilanges Lab (4) Apply Ilanges Lab filter
- Jayaraman Lab (48) Apply Jayaraman Lab filter
- Ji Lab (33) Apply Ji Lab filter
- Johnson Lab (6) Apply Johnson Lab filter
- Kainmueller Lab (19) Apply Kainmueller Lab filter
- Karpova Lab (14) Apply Karpova Lab filter
- Keleman Lab (13) Apply Keleman Lab filter
- Keller Lab (76) Apply Keller Lab filter
- Koay Lab (19) Apply Koay Lab filter
- Lavis Lab (158) Apply Lavis Lab filter
- Lee (Albert) Lab (34) Apply Lee (Albert) Lab filter
- Leonardo Lab (23) Apply Leonardo Lab filter
- Li Lab (32) Apply Li Lab filter
- Lippincott-Schwartz Lab (180) Apply Lippincott-Schwartz Lab filter
- Liu (Yin) Lab (8) Apply Liu (Yin) Lab filter
- Liu (Zhe) Lab (65) Apply Liu (Zhe) Lab filter
- Looger Lab (138) Apply Looger Lab filter
- Magee Lab (49) Apply Magee Lab filter
- Menon Lab (18) Apply Menon Lab filter
- Murphy Lab (13) Apply Murphy Lab filter
- O'Shea Lab (8) Apply O'Shea Lab filter
- Otopalik Lab (13) Apply Otopalik Lab filter
- Pachitariu Lab (52) Apply Pachitariu Lab filter
- Pastalkova Lab (19) Apply Pastalkova Lab filter
- Pavlopoulos Lab (19) Apply Pavlopoulos Lab filter
- Pedram Lab (15) Apply Pedram Lab filter
- Podgorski Lab (16) Apply Podgorski Lab filter
- Reiser Lab (55) Apply Reiser Lab filter
- Riddiford Lab (44) Apply Riddiford Lab filter
- Romani Lab (51) Apply Romani Lab filter
- Rubin Lab (149) Apply Rubin Lab filter
- Saalfeld Lab (64) Apply Saalfeld Lab filter
- Satou Lab (18) Apply Satou Lab filter
- Scheffer Lab (38) Apply Scheffer Lab filter
- Schreiter Lab (70) Apply Schreiter Lab filter
- Schulze Lab (1) Apply Schulze Lab filter
- Sgro Lab (23) Apply Sgro Lab filter
- Shroff Lab (31) Apply Shroff Lab filter
- Simpson Lab (23) Apply Simpson Lab filter
- Singer Lab (80) Apply Singer Lab filter
- Spruston Lab (98) Apply Spruston Lab filter
- Stern Lab (160) Apply Stern Lab filter
- Sternson Lab (54) Apply Sternson Lab filter
- Stringer Lab (41) Apply Stringer Lab filter
- Svoboda Lab (136) Apply Svoboda Lab filter
- Tebo Lab (35) Apply Tebo Lab filter
- Tervo Lab (9) Apply Tervo Lab filter
- Tillberg Lab (22) Apply Tillberg Lab filter
- Tjian Lab (64) Apply Tjian Lab filter
- Truman Lab (88) Apply Truman Lab filter
- Turaga Lab (53) Apply Turaga Lab filter
- Turner Lab (38) Apply Turner Lab filter
- Vale Lab (8) Apply Vale Lab filter
- Voigts Lab (4) Apply Voigts Lab filter
- Wang (Meng) Lab (29) Apply Wang (Meng) Lab filter
- Wang (Shaohe) Lab (25) Apply Wang (Shaohe) Lab filter
- Wong-Campos Lab (1) Apply Wong-Campos Lab filter
- Wu Lab (9) Apply Wu Lab filter
- Zlatic Lab (28) Apply Zlatic Lab filter
- Zuker Lab (25) Apply Zuker Lab filter
Associated Project Team
- CellMap (13) Apply CellMap filter
- COSEM (3) Apply COSEM filter
- FIB-SEM Technology (5) Apply FIB-SEM Technology filter
- Fly Descending Interneuron (12) Apply Fly Descending Interneuron filter
- Fly Functional Connectome (14) Apply Fly Functional Connectome filter
- Fly Olympiad (5) Apply Fly Olympiad filter
- FlyEM (56) Apply FlyEM filter
- FlyLight (50) Apply FlyLight filter
- GENIE (47) Apply GENIE filter
- Integrative Imaging (9) Apply Integrative Imaging filter
- Larval Olympiad (2) Apply Larval Olympiad filter
- MouseLight (18) Apply MouseLight filter
- NeuroSeq (1) Apply NeuroSeq filter
- ThalamoSeq (1) Apply ThalamoSeq filter
- Tool Translation Team (T3) (29) Apply Tool Translation Team (T3) filter
- Transcription Imaging (49) Apply Transcription Imaging filter
Publication Date
- 2026 (70) Apply 2026 filter
- 2025 (222) Apply 2025 filter
- 2024 (210) Apply 2024 filter
- 2023 (158) Apply 2023 filter
- 2022 (192) Apply 2022 filter
- 2021 (194) Apply 2021 filter
- 2020 (196) Apply 2020 filter
- 2019 (202) Apply 2019 filter
- 2018 (232) Apply 2018 filter
- 2017 (217) Apply 2017 filter
- 2016 (209) Apply 2016 filter
- 2015 (252) Apply 2015 filter
- 2014 (236) Apply 2014 filter
- 2013 (194) Apply 2013 filter
- 2012 (190) Apply 2012 filter
- 2011 (190) Apply 2011 filter
- 2010 (161) Apply 2010 filter
- 2009 (158) Apply 2009 filter
- 2008 (140) Apply 2008 filter
- 2007 (106) Apply 2007 filter
- 2006 (92) Apply 2006 filter
- 2005 (67) Apply 2005 filter
- 2004 (57) Apply 2004 filter
- 2003 (58) Apply 2003 filter
- 2002 (39) Apply 2002 filter
- 2001 (28) Apply 2001 filter
- 2000 (29) Apply 2000 filter
- 1999 (14) Apply 1999 filter
- 1998 (18) Apply 1998 filter
- 1997 (16) Apply 1997 filter
- 1996 (10) Apply 1996 filter
- 1995 (18) Apply 1995 filter
- 1994 (12) Apply 1994 filter
- 1993 (10) Apply 1993 filter
- 1992 (6) Apply 1992 filter
- 1991 (11) Apply 1991 filter
- 1990 (11) Apply 1990 filter
- 1989 (6) Apply 1989 filter
- 1988 (1) Apply 1988 filter
- 1987 (7) Apply 1987 filter
- 1986 (4) Apply 1986 filter
- 1985 (5) Apply 1985 filter
- 1984 (2) Apply 1984 filter
- 1983 (2) Apply 1983 filter
- 1982 (3) Apply 1982 filter
- 1981 (3) Apply 1981 filter
- 1980 (1) Apply 1980 filter
- 1979 (1) Apply 1979 filter
- 1976 (2) Apply 1976 filter
- 1973 (1) Apply 1973 filter
- 1970 (1) Apply 1970 filter
- 1967 (1) Apply 1967 filter
Type of Publication
4265 Publications
Showing 2101-2110 of 4265 resultsEver since the integration of Mendelian genetics into evolutionary biology in the early 20th century, evolutionary geneticists have for the most part treated genes and mutations as generic entities. However, recent observations indicate that all genes are not equal in the eyes of evolution. Evolutionarily relevant mutations tend to accumulate in hotspot genes and at specific positions within genes. Genetic evolution is constrained by gene function, the structure of genetic networks, and population biology. The genetic basis of evolution may be predictable to some extent, and further understanding of this predictability requires incorporation of the specific functions and characteristics of genes into evolutionary theory.
A nuclear gene (QCR9) encoding the 7.3-kDa subunit 9 of the mitochondrial cytochrome bc1 complex from Saccharomyces cerevisiae has been isolated from a yeast genomic library by hybridization with a degenerate oligonucleotide corresponding to nine amino acids proximal to the N terminus of purified subunit 9. QCR9 includes a 195-base pair open reading frame capable of encoding a protein of 66 amino acids and having a predicted molecular weight of 7471. The N-terminal methionine of subunit 9 is removed posttranslationally because the N-terminal sequence of the purified protein begins with serine 2. The ATG triplet corresponding to the N-terminal methionine is separated from the open reading frame by an intron. The intron is 213 base pairs long and contains previously reported 5’ donor, 3’ acceptor, and TACTAAC sequences necessary for splicing. The splice junctions, as well as the 5’ end of the message, were confirmed by isolation and sequencing of a cDNA copy of QCR9. In addition, the intron contains a nucleotide sequence in which 15 out of 18 nucleotides are identical with a sequence in the intron of COX4, the nuclear gene encoding cytochrome c oxidase subunit 4. The deduced amino acid sequence of the yeast subunit 9 is 39% identical with that of a protein of similar molecular weight from beef heart cytochrome bc1 complex. If conservative substitutions are allowed for, the two proteins are 56% similar. The predicted secondary structure of the 7.3-kDa protein revealed a single possible transmembrane helix, in which the amino acids conserved between beef heart and yeast are asymmetrically arranged along one face of the helix, implying that this domain of the protein is involved in a conserved interaction with another hydrophobic protein of the cytochrome bc1 complex. Two yeast strains, JDP1 and JDP2, were constructed in which QCR9 was deleted. Both strains grew very poorly, or not at all, on nonfermentable carbon sources and exhibited, at most, only 5% of wild-type ubiquinol-cytochrome c oxidoreductase activity. Optical spectra of mitochondrial membranes from the deletion strains revealed slightly reduced levels of cytochrome b. When JDP1 and JDP2 were complemented with a plasmid carrying QCR9, the resulting yeast grew normally on ethanol/glycerol and exhibited normal cytochrome c reductase activities and optical spectra. These results indicate that QCR9 encodes a 7.3-kDa subunit of the bc1 complex that is required for formation of a fully functional complex.(ABSTRACT TRUNCATED AT 400 WORDS)
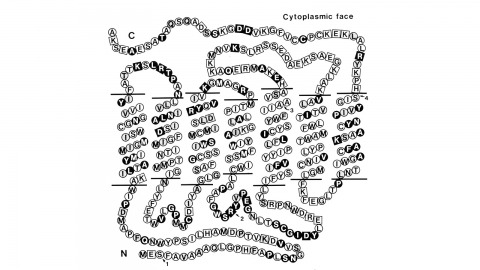
Using a novel method for detecting cross-homologous nucleic acid sequences we have isolated the gene coding for the major rhodopsin of Drosophila melanogaster and mapped it to chromosomal region 92B8-11. Comparison of cDNA and genomic DNA sequences indicates that the gene is divided into five exons. The amino acid sequence deduced from the nucleotide sequence is 373 residues long, and the polypeptide chain contains seven hydrophobic segments that appear to correspond to the seven transmembrane segments characteristic of other rhodopsins. Three regions of Drosophila rhodopsin are highly conserved with the corresponding domains of bovine rhodopsin, suggesting an important role for these polypeptide regions.
Probing chromatin structure with nucleases is a well-established method for determining the accessibility of DNA to gene regulatory proteins and measuring competency for transcription. A hallmark of many silent genes is the presence of translationally positioned nucleosomes over their promoter regions, which can be inferred by the sensitivity of the underlying DNA to nucleases, particularly micrococcal nuclease. The quality of this data is highly dependent upon the nuclear preparation, especially if the digestion products are analyzed by high-resolution detection methods such as reiterative primer extension. Here we describe a method to isolate highly purified nuclei from the budding yeast Saccharomyces cerevisiae and the use of micrococcal nuclease to map the positions of nucleosomes at the RNR3 gene. Nuclei isolated by this procedure are competent for many of the commonly used chromatin mapping and detection procedures.
Rat or mouse liver is the most frequently used tissue for mitochondrial preparations because it is readily available, easy to homogenize, and replete with mitochondria. A motor-driven Teflon and glass Potter-Elvehjem homogenizer is the best choice for homogenizing liver, but if one is not available, this tissue is soft enough that a Dounce homogenizer with a loose (A) pestle can also be used. The yield and purity of the mitochondrial preparation will be influenced by the method and speed of preparation and the age and physiological condition of the animal.
Mitochondria are complex organelles at the center of cellular metabolism, apoptosis, and signaling. They continue to be the subject of intense basic investigation to understand their composition and function, but they have also captivated the attention of clinical researchers because of the growing knowledge of the (sometimes unexpected) roles of mitochondria in human diseases and aging. A full understanding of these intriguing organelles often requires their purification from cells or tissues under specific physiological or pathological conditions. Here we provide some introductory considerations for those interested in purifying mitochondria for subsequent downstream biophysical, structural, and functional analysis.
The number of mitochondria per cell varies substantially from cell line to cell line. For example, human HeLa cells contain at least twice as many mitochondria as smaller mouse L cells. This protocol starts with a washed cell pellet of 1-2 mL derived from ∼10⁹ cells grown in culture. The cells are swollen in a hypotonic buffer and ruptured with a Dounce or Potter-Elvehjem homogenizer using a tight-fitting pestle, and mitochondria are isolated by differential centrifugation.
Chromatin structure and nucleosome positioning play a crucial role in gene expression regulation. Nucleosome positioning is often inferred by the protection of underlying DNA to nucleases. Because nucleases are excluded by plasma membranes, chromatin mapping requires isolating nuclei from cells and digesting the chromatin in situ with nucleases. The quality of this data is highly dependent on the nuclei preparation. Here we describe a method to isolate nuclei from the budding yeast Saccharomyces cerevisiae and the use of micrococcal nuclease to map the chromatin structure at the RNR3 gene. Nuclei isolated by this procedure are competent for many of the common chromatin mapping and detection procedures.
Ionotropic glutamate receptors (iGluRs) at postsynaptic terminals mediate the majority of fast excitatory neurotransmission in response to release of glutamate from the presynaptic terminal. Obtaining structural information on the molecular organization of iGluRs in their native environment, along with other signaling and scaffolding proteins in the postsynaptic density (PSD), and associated proteins on the presynaptic terminal, would enhance understanding of the molecular basis for excitatory synaptic transmission in normal and in disease states. Cryo-electron tomography (ET) studies of synaptosomes is one attractive vehicle by which to study iGluR-containing excitatory synapses. Here we describe a workflow for the preparation of glutamatergic synaptosomes for cryo-ET studies. We describe the utilization of fluorescent markers for the facile detection of the pre and postsynaptic terminals of glutamatergic synaptosomes using cryo-laser scanning confocal microscope (cryo-LSM). We further provide the details for preparation of lamellae, between ~100 to 200 nm thick, of glutamatergic synaptosomes using cryo-focused ion-beam (FIB) milling. We monitor the lamella preparation using a scanning electron microscope (SEM) and following lamella production, we identify regions for subsequent cryo-ET studies by confocal fluorescent imaging, exploiting the pre and postsynaptic fluorophores.
Targeting small-molecule fluorescent indicators using genetically encoded protein tags yields new hybrid sensors for biological imaging. Optimization of such systems requires redesign of the synthetic indicator to allow cell-specific targeting without compromising the photophysical properties or cellular performance of the small-molecule probe. We developed a bright and sensitive Ca indicator by systematically exploring the relative configuration of dye and chelator, which can be targeted using the HaloTag self-labeling tag system. Our "isomeric tuning" approach is generalizable, yielding a far-red targetable indicator to visualize Ca fluxes in the primary cilium.