Filter
Associated Lab
- Aguilera Castrejon Lab (17) Apply Aguilera Castrejon Lab filter
- Ahrens Lab (68) Apply Ahrens Lab filter
- Aso Lab (42) Apply Aso Lab filter
- Baker Lab (38) Apply Baker Lab filter
- Betzig Lab (115) Apply Betzig Lab filter
- Beyene Lab (14) Apply Beyene Lab filter
- Bock Lab (17) Apply Bock Lab filter
- Branson Lab (54) Apply Branson Lab filter
- Card Lab (43) Apply Card Lab filter
- Cardona Lab (64) Apply Cardona Lab filter
- Chklovskii Lab (13) Apply Chklovskii Lab filter
- Clapham Lab (15) Apply Clapham Lab filter
- Cui Lab (19) Apply Cui Lab filter
- Darshan Lab (12) Apply Darshan Lab filter
- Dennis Lab (1) Apply Dennis Lab filter
- Dickson Lab (46) Apply Dickson Lab filter
- Druckmann Lab (25) Apply Druckmann Lab filter
- Dudman Lab (52) Apply Dudman Lab filter
- Eddy/Rivas Lab (30) Apply Eddy/Rivas Lab filter
- Egnor Lab (11) Apply Egnor Lab filter
- Espinosa Medina Lab (20) Apply Espinosa Medina Lab filter
- Feliciano Lab (8) Apply Feliciano Lab filter
- Fetter Lab (41) Apply Fetter Lab filter
- FIB-SEM Technology (1) Apply FIB-SEM Technology filter
- Fitzgerald Lab (29) Apply Fitzgerald Lab filter
- Freeman Lab (15) Apply Freeman Lab filter
- Funke Lab (41) Apply Funke Lab filter
- Gonen Lab (91) Apply Gonen Lab filter
- Grigorieff Lab (62) Apply Grigorieff Lab filter
- Harris Lab (64) Apply Harris Lab filter
- Heberlein Lab (94) Apply Heberlein Lab filter
- Hermundstad Lab (29) Apply Hermundstad Lab filter
- Hess Lab (79) Apply Hess Lab filter
- Ilanges Lab (2) Apply Ilanges Lab filter
- Jayaraman Lab (47) Apply Jayaraman Lab filter
- Ji Lab (33) Apply Ji Lab filter
- Johnson Lab (6) Apply Johnson Lab filter
- Kainmueller Lab (19) Apply Kainmueller Lab filter
- Karpova Lab (14) Apply Karpova Lab filter
- Keleman Lab (13) Apply Keleman Lab filter
- Keller Lab (76) Apply Keller Lab filter
- Koay Lab (18) Apply Koay Lab filter
- Lavis Lab (151) Apply Lavis Lab filter
- Lee (Albert) Lab (34) Apply Lee (Albert) Lab filter
- Leonardo Lab (23) Apply Leonardo Lab filter
- Li Lab (29) Apply Li Lab filter
- Lippincott-Schwartz Lab (173) Apply Lippincott-Schwartz Lab filter
- Liu (Yin) Lab (7) Apply Liu (Yin) Lab filter
- Liu (Zhe) Lab (64) Apply Liu (Zhe) Lab filter
- Looger Lab (138) Apply Looger Lab filter
- Magee Lab (49) Apply Magee Lab filter
- Menon Lab (18) Apply Menon Lab filter
- Murphy Lab (13) Apply Murphy Lab filter
- O'Shea Lab (7) Apply O'Shea Lab filter
- Otopalik Lab (13) Apply Otopalik Lab filter
- Pachitariu Lab (49) Apply Pachitariu Lab filter
- Pastalkova Lab (18) Apply Pastalkova Lab filter
- Pavlopoulos Lab (19) Apply Pavlopoulos Lab filter
- Pedram Lab (15) Apply Pedram Lab filter
- Podgorski Lab (16) Apply Podgorski Lab filter
- Reiser Lab (52) Apply Reiser Lab filter
- Riddiford Lab (44) Apply Riddiford Lab filter
- Romani Lab (48) Apply Romani Lab filter
- Rubin Lab (146) Apply Rubin Lab filter
- Saalfeld Lab (64) Apply Saalfeld Lab filter
- Satou Lab (16) Apply Satou Lab filter
- Scheffer Lab (38) Apply Scheffer Lab filter
- Schreiter Lab (68) Apply Schreiter Lab filter
- Sgro Lab (21) Apply Sgro Lab filter
- Shroff Lab (31) Apply Shroff Lab filter
- Simpson Lab (23) Apply Simpson Lab filter
- Singer Lab (80) Apply Singer Lab filter
- Spruston Lab (94) Apply Spruston Lab filter
- Stern Lab (158) Apply Stern Lab filter
- Sternson Lab (54) Apply Sternson Lab filter
- Stringer Lab (39) Apply Stringer Lab filter
- Svoboda Lab (135) Apply Svoboda Lab filter
- Tebo Lab (33) Apply Tebo Lab filter
- Tervo Lab (9) Apply Tervo Lab filter
- Tillberg Lab (21) Apply Tillberg Lab filter
- Tjian Lab (64) Apply Tjian Lab filter
- Truman Lab (88) Apply Truman Lab filter
- Turaga Lab (52) Apply Turaga Lab filter
- Turner Lab (39) Apply Turner Lab filter
- Vale Lab (8) Apply Vale Lab filter
- Voigts Lab (3) Apply Voigts Lab filter
- Wang (Meng) Lab (23) Apply Wang (Meng) Lab filter
- Wang (Shaohe) Lab (25) Apply Wang (Shaohe) Lab filter
- Wu Lab (9) Apply Wu Lab filter
- Zlatic Lab (28) Apply Zlatic Lab filter
- Zuker Lab (25) Apply Zuker Lab filter
Associated Project Team
- CellMap (12) Apply CellMap filter
- COSEM (3) Apply COSEM filter
- FIB-SEM Technology (5) Apply FIB-SEM Technology filter
- Fly Descending Interneuron (12) Apply Fly Descending Interneuron filter
- Fly Functional Connectome (14) Apply Fly Functional Connectome filter
- Fly Olympiad (5) Apply Fly Olympiad filter
- FlyEM (56) Apply FlyEM filter
- FlyLight (50) Apply FlyLight filter
- GENIE (47) Apply GENIE filter
- Integrative Imaging (6) Apply Integrative Imaging filter
- Larval Olympiad (2) Apply Larval Olympiad filter
- MouseLight (18) Apply MouseLight filter
- NeuroSeq (1) Apply NeuroSeq filter
- ThalamoSeq (1) Apply ThalamoSeq filter
- Tool Translation Team (T3) (27) Apply Tool Translation Team (T3) filter
- Transcription Imaging (49) Apply Transcription Imaging filter
Publication Date
- 2025 (193) Apply 2025 filter
- 2024 (212) Apply 2024 filter
- 2023 (159) Apply 2023 filter
- 2022 (192) Apply 2022 filter
- 2021 (194) Apply 2021 filter
- 2020 (196) Apply 2020 filter
- 2019 (202) Apply 2019 filter
- 2018 (232) Apply 2018 filter
- 2017 (217) Apply 2017 filter
- 2016 (209) Apply 2016 filter
- 2015 (252) Apply 2015 filter
- 2014 (236) Apply 2014 filter
- 2013 (194) Apply 2013 filter
- 2012 (190) Apply 2012 filter
- 2011 (190) Apply 2011 filter
- 2010 (161) Apply 2010 filter
- 2009 (158) Apply 2009 filter
- 2008 (140) Apply 2008 filter
- 2007 (106) Apply 2007 filter
- 2006 (92) Apply 2006 filter
- 2005 (67) Apply 2005 filter
- 2004 (57) Apply 2004 filter
- 2003 (58) Apply 2003 filter
- 2002 (39) Apply 2002 filter
- 2001 (28) Apply 2001 filter
- 2000 (29) Apply 2000 filter
- 1999 (14) Apply 1999 filter
- 1998 (18) Apply 1998 filter
- 1997 (16) Apply 1997 filter
- 1996 (10) Apply 1996 filter
- 1995 (18) Apply 1995 filter
- 1994 (12) Apply 1994 filter
- 1993 (10) Apply 1993 filter
- 1992 (6) Apply 1992 filter
- 1991 (11) Apply 1991 filter
- 1990 (11) Apply 1990 filter
- 1989 (6) Apply 1989 filter
- 1988 (1) Apply 1988 filter
- 1987 (7) Apply 1987 filter
- 1986 (4) Apply 1986 filter
- 1985 (5) Apply 1985 filter
- 1984 (2) Apply 1984 filter
- 1983 (2) Apply 1983 filter
- 1982 (3) Apply 1982 filter
- 1981 (3) Apply 1981 filter
- 1980 (1) Apply 1980 filter
- 1979 (1) Apply 1979 filter
- 1976 (2) Apply 1976 filter
- 1973 (1) Apply 1973 filter
- 1970 (1) Apply 1970 filter
- 1967 (1) Apply 1967 filter
Type of Publication
4169 Publications
Showing 3491-3500 of 4169 resultsAssociative memory formation and recall in the fruit fly Drosophila melanogaster is subserved by the mushroom body (MB). Upon arrival in the MB, sensory information undergoes a profound transformation from broadly tuned and stereotyped odorant responses in the olfactory projection neuron (PN) layer to narrowly tuned and nonstereotyped responses in the Kenyon cells (KCs). Theory and experiment suggest that this transformation is implemented by random connectivity between KCs and PNs. However, this hypothesis has been challenging to test, given the difficulty of mapping synaptic connections between large numbers of brain-spanning neurons. Here, we used a recent whole-brain electron microscopy volume of the adult fruit fly to map PN-to-KC connectivity at synaptic resolution. The PN-KC connectome revealed unexpected structure, with preponderantly food-responsive PN types converging at above-chance levels on downstream KCs. Axons of the overconvergent PN types tended to arborize near one another in the MB main calyx, making local KC dendrites more likely to receive input from those types. Overconvergent PN types preferentially co-arborize and connect with dendrites of αβ and α'β' KC subtypes. Computational simulation of the observed network showed degraded discrimination performance compared with a random network, except when all signal flowed through the overconvergent, primarily food-responsive PN types. Additional theory and experiment will be needed to fully characterize the impact of the observed non-random network structure on associative memory formation and recall.
The organization of synaptic connectivity within a neuronal circuit is a prime determinant of circuit function. We performed a comprehensive fine-scale circuit mapping of hippocampal regions (CA3-CA1) using the newly developed synapse labeling method, mGRASP. This mapping revealed spatially nonuniform and clustered synaptic connectivity patterns. Furthermore, synaptic clustering was enhanced between groups of neurons that shared a similar developmental/migration time window, suggesting a mechanism for establishing the spatial structure of synaptic connectivity. Such connectivity patterns are thought to effectively engage active dendritic processing and storage mechanisms, thereby potentially enhancing neuronal feature selectivity.
Sequence-specific DNA-binding activators, key regulators of gene expression, stimulate transcription in part by targeting the core promoter recognition TFIID complex and aiding in its recruitment to promoter DNA. Although it has been established that activators can interact with multiple components of TFIID, it is unknown whether common or distinct surfaces within TFIID are targeted by activators and what changes if any in the structure of TFIID may occur upon binding activators. As a first step toward structurally dissecting activator/TFIID interactions, we determined the three-dimensional structures of TFIID bound to three distinct activators (i.e., the tumor suppressor p53 protein, glutamine-rich Sp1 and the oncoprotein c-Jun) and compared their structures as determined by electron microscopy and single-particle reconstruction. By a combination of EM and biochemical mapping analysis, our results uncover distinct contact regions within TFIID bound by each activator. Unlike the coactivator CRSP/Mediator complex that undergoes drastic and global structural changes upon activator binding, instead, a rather confined set of local conserved structural changes were observed when each activator binds holo-TFIID. These results suggest that activator contact may induce unique structural features of TFIID, thus providing nanoscale information on activator-dependent TFIID assembly and transcription initiation.
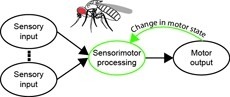
Sensorimotor integration is a field rich in theory backed by a large body of psychophysical evidence. Relating the underlying neural circuitry to these theories has, however, been more challenging. With a wide array of complex behaviors coordinated by their small brains, insects provide powerful model systems to study key features of sensorimotor integration at a mechanistic level. Insect neural circuits perform both hard-wired and learned sensorimotor transformations. They modulate their neural processing based on both internal variables, such as the animal’s behavioral state, and external ones, such as the time of day. Here we present some studies using insect model systems that have produced insights, at the level of individual neurons, about sensorimotor integration and the various ways in which it can be modified by context.
The neural underpinnings of sensorimotor integration are best studied in the context of well-characterized behavior. A rich trove of Drosophila behavioral genetics research offers a variety of well-studied behaviors and candidate brain regions that can form the bases of such studies. The development of tools to perform in vivo physiology from the Drosophila brain has made it possible to monitor activity in defined neurons in response to sensory stimuli. More recently still, it has become possible to perform recordings from identified neurons in the brain of head-fixed flies during walking or flight behaviors. In this chapter, we discuss how experiments that simultaneously monitor behavior and physiology in Drosophila can be combined with other techniques to produce testable models of sensorimotor circuit function.
Cognition encompasses a range of higher-order mental processes, such as attention, working memory, and model-based decision-making. These processes are thought to involve the dynamic interaction of multiple central brain regions. A mechanistic understanding of such computations requires not only monitoring and manipulating specific neural populations during behavior, but also knowing the connectivity of the underlying circuitry. These goals are experimentally challenging in mammals, but are feasible in numerically simpler insect brains. In Drosophila melanogaster in particular, genetic tools enable precisely targeted physiology and optogenetics in actively behaving animals. In this article we discuss how these advantages are increasingly being leveraged to study abstract neural representations and sensorimotor computations that may be relevant for cognition in both insects and mammals.
Cortical neurons exhibit temporally irregular spiking patterns and heterogeneous firing rates. These features arise in model circuits operating in a 'fluctuation-driven regime', in which fluctuations in membrane potentials emerge from the network dynamics. However, it is still debated whether the cortex operates in such a regime. We evaluated the fluctuation-driven hypothesis by analyzing spiking and sub-threshold membrane potentials of neurons in the frontal cortex of mice performing a decision-making task. We showed that while standard fluctuation-driven models successfully account for spiking statistics, they fall short in capturing the heterogeneity in sub-threshold activity. This limitation is an inevitable outcome of bombarding single-compartment neurons with a large number of pre-synaptic inputs, thereby clamping the voltage of all neurons to more or less the same average voltage. To address this, we effectively incorporated dendritic morphology into the standard models. Inclusion of dendritic morphology in the neuronal models increased neuronal selectivity and reduced error trials, suggesting a functional role for dendrites during decision-making. Our work suggests that, during decision-making, cortical neurons in high-order cortical areas operate in a fluctuation-driven regime.
The atomic structure of the infectious, protease-resistant, β-sheet-rich and fibrillar mammalian prion remains unknown. Through the cryo-EM method MicroED, we reveal the sub-ångström-resolution structure of a protofibril formed by a wild-type segment from the β2-α2 loop of the bank vole prion protein. The structure of this protofibril reveals a stabilizing network of hydrogen bonds that link polar zippers within a sheet, producing motifs we have named 'polar clasps'.
Protein kinase A (PKA) plays multiple roles in neurons. The localization and specificity of PKA are largely controlled by A-kinase anchoring proteins (AKAPs). However, the dynamics of PKA in neurons and the roles of specific AKAPs are poorly understood. We imaged the distribution of type II PKA in hippocampal and cortical layer 2/3 pyramidal neurons in vitro and in vivo. PKA was concentrated in dendritic shafts compared to the soma, axons, and dendritic spines. This spatial distribution was imposed by the microtubule-binding protein MAP2, indicating that MAP2 is the dominant AKAP in neurons. Following cAMP elevation, catalytic subunits dissociated from the MAP2-tethered regulatory subunits and rapidly became enriched in nearby spines. The spatial gradient of type II PKA between dendritic shafts and spines was critical for the regulation of synaptic strength and long-term potentiation. Therefore, the localization and activity-dependent translocation of type II PKA are important determinants of PKA function.