Filter
Associated Lab
- Aguilera Castrejon Lab (19) Apply Aguilera Castrejon Lab filter
- Ahrens Lab (73) Apply Ahrens Lab filter
- Aso Lab (42) Apply Aso Lab filter
- Baker Lab (38) Apply Baker Lab filter
- Betzig Lab (116) Apply Betzig Lab filter
- Beyene Lab (15) Apply Beyene Lab filter
- Bock Lab (17) Apply Bock Lab filter
- Branson Lab (55) Apply Branson Lab filter
- Card Lab (43) Apply Card Lab filter
- Cardona Lab (64) Apply Cardona Lab filter
- Chklovskii Lab (13) Apply Chklovskii Lab filter
- Clapham Lab (16) Apply Clapham Lab filter
- Cui Lab (19) Apply Cui Lab filter
- Darshan Lab (12) Apply Darshan Lab filter
- Dennis Lab (3) Apply Dennis Lab filter
- Dickson Lab (46) Apply Dickson Lab filter
- Druckmann Lab (25) Apply Druckmann Lab filter
- Dudman Lab (56) Apply Dudman Lab filter
- Eddy/Rivas Lab (30) Apply Eddy/Rivas Lab filter
- Egnor Lab (11) Apply Egnor Lab filter
- Espinosa Medina Lab (23) Apply Espinosa Medina Lab filter
- Feliciano Lab (12) Apply Feliciano Lab filter
- Fetter Lab (41) Apply Fetter Lab filter
- FIB-SEM Technology (1) Apply FIB-SEM Technology filter
- Fitzgerald Lab (30) Apply Fitzgerald Lab filter
- Freeman Lab (15) Apply Freeman Lab filter
- Funke Lab (46) Apply Funke Lab filter
- Gonen Lab (91) Apply Gonen Lab filter
- Grigorieff Lab (62) Apply Grigorieff Lab filter
- Harris Lab (65) Apply Harris Lab filter
- Heberlein Lab (94) Apply Heberlein Lab filter
- Hermundstad Lab (30) Apply Hermundstad Lab filter
- Hess Lab (80) Apply Hess Lab filter
- Ilanges Lab (4) Apply Ilanges Lab filter
- Jayaraman Lab (48) Apply Jayaraman Lab filter
- Ji Lab (33) Apply Ji Lab filter
- Johnson Lab (6) Apply Johnson Lab filter
- Kainmueller Lab (19) Apply Kainmueller Lab filter
- Karpova Lab (14) Apply Karpova Lab filter
- Keleman Lab (13) Apply Keleman Lab filter
- Keller Lab (76) Apply Keller Lab filter
- Koay Lab (19) Apply Koay Lab filter
- Lavis Lab (158) Apply Lavis Lab filter
- Lee (Albert) Lab (34) Apply Lee (Albert) Lab filter
- Leonardo Lab (23) Apply Leonardo Lab filter
- Li Lab (32) Apply Li Lab filter
- Lippincott-Schwartz Lab (180) Apply Lippincott-Schwartz Lab filter
- Liu (Yin) Lab (8) Apply Liu (Yin) Lab filter
- Liu (Zhe) Lab (65) Apply Liu (Zhe) Lab filter
- Looger Lab (138) Apply Looger Lab filter
- Magee Lab (49) Apply Magee Lab filter
- Menon Lab (18) Apply Menon Lab filter
- Murphy Lab (13) Apply Murphy Lab filter
- O'Shea Lab (8) Apply O'Shea Lab filter
- Otopalik Lab (13) Apply Otopalik Lab filter
- Pachitariu Lab (52) Apply Pachitariu Lab filter
- Pastalkova Lab (19) Apply Pastalkova Lab filter
- Pavlopoulos Lab (19) Apply Pavlopoulos Lab filter
- Pedram Lab (15) Apply Pedram Lab filter
- Podgorski Lab (16) Apply Podgorski Lab filter
- Reiser Lab (55) Apply Reiser Lab filter
- Riddiford Lab (44) Apply Riddiford Lab filter
- Romani Lab (51) Apply Romani Lab filter
- Rubin Lab (149) Apply Rubin Lab filter
- Saalfeld Lab (64) Apply Saalfeld Lab filter
- Satou Lab (18) Apply Satou Lab filter
- Scheffer Lab (38) Apply Scheffer Lab filter
- Schreiter Lab (70) Apply Schreiter Lab filter
- Schulze Lab (1) Apply Schulze Lab filter
- Sgro Lab (23) Apply Sgro Lab filter
- Shroff Lab (31) Apply Shroff Lab filter
- Simpson Lab (23) Apply Simpson Lab filter
- Singer Lab (80) Apply Singer Lab filter
- Spruston Lab (98) Apply Spruston Lab filter
- Stern Lab (160) Apply Stern Lab filter
- Sternson Lab (54) Apply Sternson Lab filter
- Stringer Lab (41) Apply Stringer Lab filter
- Svoboda Lab (136) Apply Svoboda Lab filter
- Tebo Lab (35) Apply Tebo Lab filter
- Tervo Lab (9) Apply Tervo Lab filter
- Tillberg Lab (22) Apply Tillberg Lab filter
- Tjian Lab (64) Apply Tjian Lab filter
- Truman Lab (88) Apply Truman Lab filter
- Turaga Lab (53) Apply Turaga Lab filter
- Turner Lab (38) Apply Turner Lab filter
- Vale Lab (8) Apply Vale Lab filter
- Voigts Lab (4) Apply Voigts Lab filter
- Wang (Meng) Lab (29) Apply Wang (Meng) Lab filter
- Wang (Shaohe) Lab (25) Apply Wang (Shaohe) Lab filter
- Wong-Campos Lab (1) Apply Wong-Campos Lab filter
- Wu Lab (9) Apply Wu Lab filter
- Zlatic Lab (28) Apply Zlatic Lab filter
- Zuker Lab (25) Apply Zuker Lab filter
Associated Project Team
- CellMap (13) Apply CellMap filter
- COSEM (3) Apply COSEM filter
- FIB-SEM Technology (5) Apply FIB-SEM Technology filter
- Fly Descending Interneuron (12) Apply Fly Descending Interneuron filter
- Fly Functional Connectome (14) Apply Fly Functional Connectome filter
- Fly Olympiad (5) Apply Fly Olympiad filter
- FlyEM (56) Apply FlyEM filter
- FlyLight (50) Apply FlyLight filter
- GENIE (47) Apply GENIE filter
- Integrative Imaging (9) Apply Integrative Imaging filter
- Larval Olympiad (2) Apply Larval Olympiad filter
- MouseLight (18) Apply MouseLight filter
- NeuroSeq (1) Apply NeuroSeq filter
- ThalamoSeq (1) Apply ThalamoSeq filter
- Tool Translation Team (T3) (29) Apply Tool Translation Team (T3) filter
- Transcription Imaging (49) Apply Transcription Imaging filter
Publication Date
- 2026 (70) Apply 2026 filter
- 2025 (222) Apply 2025 filter
- 2024 (210) Apply 2024 filter
- 2023 (158) Apply 2023 filter
- 2022 (192) Apply 2022 filter
- 2021 (194) Apply 2021 filter
- 2020 (196) Apply 2020 filter
- 2019 (202) Apply 2019 filter
- 2018 (232) Apply 2018 filter
- 2017 (217) Apply 2017 filter
- 2016 (209) Apply 2016 filter
- 2015 (252) Apply 2015 filter
- 2014 (236) Apply 2014 filter
- 2013 (194) Apply 2013 filter
- 2012 (190) Apply 2012 filter
- 2011 (190) Apply 2011 filter
- 2010 (161) Apply 2010 filter
- 2009 (158) Apply 2009 filter
- 2008 (140) Apply 2008 filter
- 2007 (106) Apply 2007 filter
- 2006 (92) Apply 2006 filter
- 2005 (67) Apply 2005 filter
- 2004 (57) Apply 2004 filter
- 2003 (58) Apply 2003 filter
- 2002 (39) Apply 2002 filter
- 2001 (28) Apply 2001 filter
- 2000 (29) Apply 2000 filter
- 1999 (14) Apply 1999 filter
- 1998 (18) Apply 1998 filter
- 1997 (16) Apply 1997 filter
- 1996 (10) Apply 1996 filter
- 1995 (18) Apply 1995 filter
- 1994 (12) Apply 1994 filter
- 1993 (10) Apply 1993 filter
- 1992 (6) Apply 1992 filter
- 1991 (11) Apply 1991 filter
- 1990 (11) Apply 1990 filter
- 1989 (6) Apply 1989 filter
- 1988 (1) Apply 1988 filter
- 1987 (7) Apply 1987 filter
- 1986 (4) Apply 1986 filter
- 1985 (5) Apply 1985 filter
- 1984 (2) Apply 1984 filter
- 1983 (2) Apply 1983 filter
- 1982 (3) Apply 1982 filter
- 1981 (3) Apply 1981 filter
- 1980 (1) Apply 1980 filter
- 1979 (1) Apply 1979 filter
- 1976 (2) Apply 1976 filter
- 1973 (1) Apply 1973 filter
- 1970 (1) Apply 1970 filter
- 1967 (1) Apply 1967 filter
Type of Publication
4265 Publications
Showing 3831-3840 of 4265 resultsRodents move their whiskers to locate objects in space. Here we used psychophysical methods to show that head-fixed mice can localize objects along the axis of a single whisker, the radial dimension, with one-millimeter precision. High-speed videography allowed us to estimate the forces and bending moments at the base of the whisker, which underlie radial distance measurement. Mice judged radial object location based on multiple touches. Both the number of touches (1-17) and the forces exerted by the pole on the whisker (up to 573 μN; typical peak amplitude, 100 μN) varied greatly across trials. We manipulated the bending moment and lateral force pressing the whisker against the sides of the follicle and the axial force pushing the whisker into the follicle by varying the compliance of the object during behavior. The behavioral responses suggest that mice use multiple variables (bending moment, axial force, lateral force) to extract radial object localization. Characterization of whisker mechanics revealed that whisker bending stiffness decreases gradually with distance from the face over five orders of magnitude. As a result, the relative amplitudes of different stress variables change dramatically with radial object distance. Our data suggest that mice use distance-dependent whisker mechanics to estimate radial object location using an algorithm that does not rely on precise control of whisking, is robust to variability in whisker forces, and is independent of object compliance and object movement. More generally, our data imply that mice can measure the amplitudes of forces in the sensory follicles for tactile sensation.
Several types of retinal interneurons exhibit spikes but lack axons. One such neuron is the AII amacrine cell, in which spikes recorded at the soma exhibit small amplitudes (<10 mV) and broad time courses (>5 ms). Here, we used electrophysiological recordings and computational analysis to examine the mechanisms underlying this atypical spiking. We found that somatic spikes likely represent large, brief action potential-like events initiated in a single, electrotonically distal dendritic compartment. In this same compartment, spiking undergoes slow modulation, likely by an M-type K conductance. The structural correlate of this compartment is a thin neurite that extends from the primary dendritic tree: local application of TTX to this neurite, or excision of it, eliminates spiking. Thus, the physiology of the axonless AII is much more complex than would be anticipated from morphological descriptions and somatic recordings; in particular, the AII possesses a single dendritic structure that controls its firing pattern.
Summary Precise regulation of microtubule (MT) dynamics is increasingly recognized as a critical determinant of axon regeneration. In contrast to developing neurons, mature axons exhibit noncentrosomal microtubule nucleation. The factors regulating noncentrosomal MT architecture in axon regeneration remain poorly understood. We report that PTRN-1, the C. elegans member of the Patronin/Nezha/calmodulin- and spectrin-associated protein (CAMSAP) family of microtubule minus-end-binding proteins, is critical for efficient axon regeneration in vivo. ptrn-1-null mutants display generally normal developmental axon outgrowth but significantly impaired regenerative regrowth after laser axotomy. Unexpectedly, mature axons in ptrn-1 mutants display elevated numbers of dynamic axonal MTs before and after injury, suggesting that PTRN-1 inhibits MT dynamics. The CKK domain of PTRN-1 is necessary and sufficient for its functions in axon regeneration and MT dynamics and appears to stabilize MTs independent of minus-end localization. Whereas in developing neurons, PTRN-1 inhibits activity of the DLK-1 mitogen-activated protein kinase (MAPK) cascade, we find that, in regeneration, PTRN-1 and DLK-1 function together to promote axonal regrowth.
Large scientific projects in genomics and astronomy are influential not because they answer any single question but because they enable investigation of continuously arising new questions from the same data-rich sources. Advances in automated mapping of the brain's synaptic connections (connectomics) suggest that the complicated circuits underlying brain function are ripe for analysis. We discuss benefits of mapping a mouse brain at the level of synapses.
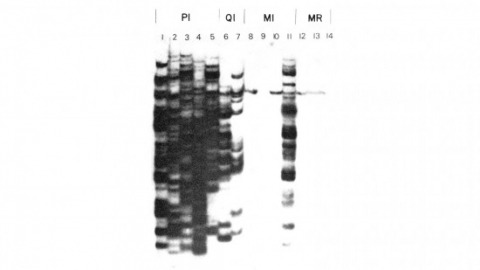
We have shown previously that four of five white mutant alleles arising in P-M dysgenic hybrids result from the insertion of strongly homologous DNA sequence elements. We have named these P elements. We report that P elements are present in 30-50 copies per haploid genome in all P strains examined and apparently are missing entirely from all M strains examined, with one exception. Furthermore, members of the P family apparently transpose frequently in P-M dysgenic hybrids; chromosomes descendant from P-M dysgenic hybrids frequently show newly acquired P elements. Finally, the strain-specific breakpoint hotspots for the rearrangement of the pi 2 P X chromosome occurring in P-M dysgenic hybrids are apparently sites of residence of P elements. These observations strongly support the P factor hypothesis for the mechanistic basis of P-M hybrid dysgenesis.
Many insect developmental color changes are known to be regulated by both ecdysone and juvenile hormone. Yet the molecular mechanisms underlying this regulation have not been well understood. This review highlights the hormonal mechanisms involved in the regulation of two key enzymes [dopa decarboxylase (DDC) and phenoloxidase] necessary for insect cuticular melanization, and the molecular action of 20-hydroxyecdysone on various transcription factors leading to DDC expression at the end of a larval molt in Manduca sexta. In addition, the ecdysone cascade found in M. sexta is compared with that of other organisms.
In Drosophila dosage compensation increases the rate of transcription of the male's X chromosome and depends on four autosomal male-specific lethal genes. We have cloned the msl-2 gene and shown that MSL-2 protein is co-localized with the other three MSL proteins at hundreds of sites along the male polytene X chromosome and that this binding requires the other three MSL proteins. msl-2 encodes a protein with a putative DNA-binding domain: the RING finger. MSL-2 protein is not produced in females and sequences in both the 5' and 3' UTRs are important for this sex-specific regulation. Furthermore, msl-2 pre-mRNA is alternatively spliced in a Sex-lethal-dependent fashion in its 5' UTR.
mTOR is a serine/threonine kinase and a master regulator of cell growth and proliferation. Raptor, a scaffolding protein that recruits substrates to mTOR complex 1 (mTORC1), is known to be phosphorylated during mitosis, but the significance of this phosphorylation remains largely unknown. Here we show that raptor expression and mTORC1 activity are dramatically reduced in cells arrested in mitosis. Expression of a non-phosphorylatable raptor mutant reactivates mTORC1 and significantly reduces cytotoxicity of the mitotic poison Taxol. This effect is mediated via degradation of PDCD4, a tumor suppressor protein that inhibits eIF4A activity and is negatively regulated by the mTORC1/S6K pathway. Moreover, pharmacological inhibition of eIF4A is able to enhance the effects of Taxol and restore sensitivity in Taxol-resistant cancer cells. These findings indicate that the mTORC1/S6K/PDCD4/eIF4A axis has a pivotal role in the death versus slippage decision during mitotic arrest and may be exploited clinically to treat tumors resistant to anti-mitotic agents.
Mucin domains are densely O-glycosylated modular protein domains that are found in a wide variety of cell surface and secreted proteins. Mucin-domain glycoproteins are known to be key players in a host of human diseases, especially cancer, wherein mucin expression and glycosylation patterns are altered. Mucin biology has been difficult to study at the molecular level, in part, because methods to manipulate and structurally characterize mucin domains are lacking. Here, we demonstrate that secreted protease of C1 esterase inhibitor (StcE), a bacterial protease from Escherichia coli, cleaves mucin domains by recognizing a discrete peptide- and glycan-based motif. We exploited StcE's unique properties to improve sequence coverage, glycosite mapping, and glycoform analysis of recombinant human mucins by mass spectrometry. We also found that StcE digests cancer-associated mucins from cultured cells and from ascites fluid derived from patients with ovarian cancer. Finally, using StcE, we discovered that sialic acid-binding Ig-type lectin-7 (Siglec-7), a glycoimmune checkpoint receptor, selectively binds sialomucins as biological ligands, whereas the related receptor Siglec-9 does not. Mucin-selective proteolysis, as exemplified by StcE, is therefore a powerful tool for the study of mucin domain structure and function.
Connectomics has focused primarily on the mapping of synaptic links in the brain; yet it is well established that extrasynaptic volume transmission, especially via monoamines and neuropeptides, is also critical to brain function and occurs primarily outside the synaptic connectome. We have mapped the putative monoamine connections, as well as a subset of neuropeptide connections, in C. elegans based on new and published gene expression data. The monoamine and neuropeptide networks exhibit distinct topological properties, with the monoamine network displaying a highly disassortative star-like structure with a rich-club of interconnected broadcasting hubs, and the neuropeptide network showing a more recurrent, highly clustered topology. Despite the low degree of overlap between the extrasynaptic (or wireless) and synaptic (or wired) connectomes, we find highly significant multilink motifs of interaction, pinpointing locations in the network where aminergic and neuropeptide signalling modulate synaptic activity. Thus, the C. elegans connectome can be mapped as a multiplex network with synaptic, gap junction, and neuromodulator layers representing alternative modes of interaction between neurons. This provides a new topological plan for understanding how aminergic and peptidergic modulation of behaviour is achieved by specific motifs and loci of integration between hard-wired synaptic or junctional circuits and extrasynaptic signals wirelessly broadcast from a small number of modulatory neurons.