Filter
Associated Lab
- Aguilera Castrejon Lab (19) Apply Aguilera Castrejon Lab filter
- Ahrens Lab (73) Apply Ahrens Lab filter
- Aso Lab (42) Apply Aso Lab filter
- Baker Lab (38) Apply Baker Lab filter
- Betzig Lab (116) Apply Betzig Lab filter
- Beyene Lab (15) Apply Beyene Lab filter
- Bock Lab (17) Apply Bock Lab filter
- Branson Lab (55) Apply Branson Lab filter
- Card Lab (43) Apply Card Lab filter
- Cardona Lab (64) Apply Cardona Lab filter
- Chklovskii Lab (13) Apply Chklovskii Lab filter
- Clapham Lab (16) Apply Clapham Lab filter
- Cui Lab (19) Apply Cui Lab filter
- Darshan Lab (12) Apply Darshan Lab filter
- Dennis Lab (3) Apply Dennis Lab filter
- Dickson Lab (46) Apply Dickson Lab filter
- Druckmann Lab (25) Apply Druckmann Lab filter
- Dudman Lab (56) Apply Dudman Lab filter
- Eddy/Rivas Lab (30) Apply Eddy/Rivas Lab filter
- Egnor Lab (11) Apply Egnor Lab filter
- Espinosa Medina Lab (23) Apply Espinosa Medina Lab filter
- Feliciano Lab (12) Apply Feliciano Lab filter
- Fetter Lab (41) Apply Fetter Lab filter
- FIB-SEM Technology (1) Apply FIB-SEM Technology filter
- Fitzgerald Lab (30) Apply Fitzgerald Lab filter
- Freeman Lab (15) Apply Freeman Lab filter
- Funke Lab (46) Apply Funke Lab filter
- Gonen Lab (91) Apply Gonen Lab filter
- Grigorieff Lab (62) Apply Grigorieff Lab filter
- Harris Lab (65) Apply Harris Lab filter
- Heberlein Lab (94) Apply Heberlein Lab filter
- Hermundstad Lab (30) Apply Hermundstad Lab filter
- Hess Lab (80) Apply Hess Lab filter
- Ilanges Lab (4) Apply Ilanges Lab filter
- Jayaraman Lab (48) Apply Jayaraman Lab filter
- Ji Lab (33) Apply Ji Lab filter
- Johnson Lab (6) Apply Johnson Lab filter
- Kainmueller Lab (19) Apply Kainmueller Lab filter
- Karpova Lab (15) Apply Karpova Lab filter
- Keleman Lab (13) Apply Keleman Lab filter
- Keller Lab (76) Apply Keller Lab filter
- Koay Lab (20) Apply Koay Lab filter
- Lavis Lab (158) Apply Lavis Lab filter
- Lee (Albert) Lab (34) Apply Lee (Albert) Lab filter
- Leonardo Lab (23) Apply Leonardo Lab filter
- Li Lab (32) Apply Li Lab filter
- Lippincott-Schwartz Lab (180) Apply Lippincott-Schwartz Lab filter
- Liu (Yin) Lab (8) Apply Liu (Yin) Lab filter
- Liu (Zhe) Lab (65) Apply Liu (Zhe) Lab filter
- Looger Lab (138) Apply Looger Lab filter
- Magee Lab (49) Apply Magee Lab filter
- Menon Lab (18) Apply Menon Lab filter
- Murphy Lab (13) Apply Murphy Lab filter
- O'Shea Lab (8) Apply O'Shea Lab filter
- Otopalik Lab (13) Apply Otopalik Lab filter
- Pachitariu Lab (54) Apply Pachitariu Lab filter
- Pastalkova Lab (19) Apply Pastalkova Lab filter
- Pavlopoulos Lab (19) Apply Pavlopoulos Lab filter
- Pedram Lab (15) Apply Pedram Lab filter
- Podgorski Lab (16) Apply Podgorski Lab filter
- Reiser Lab (55) Apply Reiser Lab filter
- Riddiford Lab (44) Apply Riddiford Lab filter
- Romani Lab (51) Apply Romani Lab filter
- Rubin Lab (149) Apply Rubin Lab filter
- Saalfeld Lab (64) Apply Saalfeld Lab filter
- Satou Lab (18) Apply Satou Lab filter
- Scheffer Lab (38) Apply Scheffer Lab filter
- Schreiter Lab (70) Apply Schreiter Lab filter
- Schulze Lab (1) Apply Schulze Lab filter
- Sgro Lab (23) Apply Sgro Lab filter
- Shroff Lab (31) Apply Shroff Lab filter
- Simpson Lab (23) Apply Simpson Lab filter
- Singer Lab (80) Apply Singer Lab filter
- Spruston Lab (98) Apply Spruston Lab filter
- Stern Lab (160) Apply Stern Lab filter
- Sternson Lab (54) Apply Sternson Lab filter
- Stringer Lab (43) Apply Stringer Lab filter
- Svoboda Lab (136) Apply Svoboda Lab filter
- Tebo Lab (35) Apply Tebo Lab filter
- Tervo Lab (10) Apply Tervo Lab filter
- Tillberg Lab (22) Apply Tillberg Lab filter
- Tjian Lab (64) Apply Tjian Lab filter
- Truman Lab (88) Apply Truman Lab filter
- Turaga Lab (53) Apply Turaga Lab filter
- Turner Lab (38) Apply Turner Lab filter
- Vale Lab (8) Apply Vale Lab filter
- Voigts Lab (4) Apply Voigts Lab filter
- Wang (Meng) Lab (29) Apply Wang (Meng) Lab filter
- Wang (Shaohe) Lab (25) Apply Wang (Shaohe) Lab filter
- Wong-Campos Lab (1) Apply Wong-Campos Lab filter
- Wu Lab (9) Apply Wu Lab filter
- Zlatic Lab (28) Apply Zlatic Lab filter
- Zuker Lab (25) Apply Zuker Lab filter
Associated Project Team
- CellMap (13) Apply CellMap filter
- COSEM (3) Apply COSEM filter
- FIB-SEM Technology (5) Apply FIB-SEM Technology filter
- Fly Descending Interneuron (12) Apply Fly Descending Interneuron filter
- Fly Functional Connectome (14) Apply Fly Functional Connectome filter
- Fly Olympiad (5) Apply Fly Olympiad filter
- FlyEM (56) Apply FlyEM filter
- FlyLight (50) Apply FlyLight filter
- GENIE (47) Apply GENIE filter
- Integrative Imaging (9) Apply Integrative Imaging filter
- Larval Olympiad (2) Apply Larval Olympiad filter
- MouseLight (18) Apply MouseLight filter
- NeuroSeq (1) Apply NeuroSeq filter
- ThalamoSeq (1) Apply ThalamoSeq filter
- Tool Translation Team (T3) (29) Apply Tool Translation Team (T3) filter
- Transcription Imaging (49) Apply Transcription Imaging filter
Publication Date
- 2026 (74) Apply 2026 filter
- 2025 (222) Apply 2025 filter
- 2024 (210) Apply 2024 filter
- 2023 (158) Apply 2023 filter
- 2022 (192) Apply 2022 filter
- 2021 (194) Apply 2021 filter
- 2020 (196) Apply 2020 filter
- 2019 (202) Apply 2019 filter
- 2018 (232) Apply 2018 filter
- 2017 (217) Apply 2017 filter
- 2016 (209) Apply 2016 filter
- 2015 (252) Apply 2015 filter
- 2014 (236) Apply 2014 filter
- 2013 (194) Apply 2013 filter
- 2012 (190) Apply 2012 filter
- 2011 (190) Apply 2011 filter
- 2010 (161) Apply 2010 filter
- 2009 (158) Apply 2009 filter
- 2008 (140) Apply 2008 filter
- 2007 (106) Apply 2007 filter
- 2006 (92) Apply 2006 filter
- 2005 (67) Apply 2005 filter
- 2004 (57) Apply 2004 filter
- 2003 (58) Apply 2003 filter
- 2002 (39) Apply 2002 filter
- 2001 (28) Apply 2001 filter
- 2000 (29) Apply 2000 filter
- 1999 (14) Apply 1999 filter
- 1998 (18) Apply 1998 filter
- 1997 (16) Apply 1997 filter
- 1996 (10) Apply 1996 filter
- 1995 (18) Apply 1995 filter
- 1994 (12) Apply 1994 filter
- 1993 (10) Apply 1993 filter
- 1992 (6) Apply 1992 filter
- 1991 (11) Apply 1991 filter
- 1990 (11) Apply 1990 filter
- 1989 (6) Apply 1989 filter
- 1988 (1) Apply 1988 filter
- 1987 (7) Apply 1987 filter
- 1986 (4) Apply 1986 filter
- 1985 (5) Apply 1985 filter
- 1984 (2) Apply 1984 filter
- 1983 (2) Apply 1983 filter
- 1982 (3) Apply 1982 filter
- 1981 (3) Apply 1981 filter
- 1980 (1) Apply 1980 filter
- 1979 (1) Apply 1979 filter
- 1976 (2) Apply 1976 filter
- 1973 (1) Apply 1973 filter
- 1970 (1) Apply 1970 filter
- 1967 (1) Apply 1967 filter
Type of Publication
4269 Publications
Showing 871-880 of 4269 resultsProprioceptive sensory signals inform the CNS of the consequences of motor acts, but effective motor planning involves internal neural systems capable of anticipating actual sensory feedback. Just where and how predictive systems exert their influence remains poorly understood. We explored the possibility that spinocerebellar neurons that convey proprioceptive sensory information also integrate information from cortical command systems. Analysis of the circuitry and physiology of identified dorsal spinocerebellar tract neurons in mouse spinal cord revealed distinct populations of Clarke’s column neurons that received direct excitatory and/or indirect inhibitory inputs from descending corticospinal axons. The convergence of these descending inhibitory and excitatory inputs to Clarke’s column neurons established local spinal circuits with the capacity to mark or modulate incoming proprioceptive input. Together, our genetic, anatomical and physiological results indicate that Clarke’s column spinocerebellar neurons nucleate local spinal corollary circuits that are relevant to motor planning and evaluation.
Brain function relies on specificity of synaptic connectivity patterns among different classes of neurons. Yet, the substrates of specificity in complex neuropil remain largely unknown. We search for imprints of specificity in the layout of axonal and dendritic arbors from the rat neocortex. An analysis of 3D reconstructions of pairs consisting of pyramidal cells (PCs) and GABAergic interneurons (GIs) revealed that the layout of GI axons is specific. This specificity is manifested in a relatively high tortuosity, small branch length of these axons, and correlations of their trajectories with the positions of postsynaptic neuron dendrites. Axons of PCs show no such specificity, usually taking a relatively straight course through neuropil. However, wiring patterns among PCs hold a large potential for circuit remodeling and specificity through growth and retraction of dendritic spines. Our results define distinct class-specific rules in establishing synaptic connectivity, which could be crucial in formulating a canonical cortical circuit.
Neural circuits carry out complex computations that allow animals to evaluate food, select mates, move toward attractive stimuli, and move away from threats. In insects, the subesophageal zone (SEZ) is a brain region that receives gustatory, pheromonal, and mechanosensory inputs and contributes to the control of diverse behaviors, including feeding, grooming, and locomotion. Despite its importance in sensorimotor transformations, the study of SEZ circuits has been hindered by limited knowledge of the underlying diversity of SEZ neurons. Here, we generate a collection of split-GAL4 lines that provides precise genetic targeting of 138 different SEZ cell types in adult , comprising approximately one third of all SEZ neurons. We characterize the single cell anatomy of these neurons and find that they cluster by morphology into six supergroups that organize the SEZ into discrete anatomical domains. We find that the majority of local SEZ interneurons are not classically polarized, suggesting rich local processing, whereas SEZ projection neurons tend to be classically polarized, conveying information to a limited number of higher brain regions. This study provides insight into the anatomical organization of the SEZ and generates resources that will facilitate further study of SEZ neurons and their contributions to sensory processing and behavior.
The definition of neuronal type and how it relates to the transcriptome are open questions. Drosophila olfactory projection neurons (PNs) are among the best-characterized neuronal types: different PN classes target dendrites to distinct olfactory glomeruli, while PNs of the same class exhibit indistinguishable anatomical and physiological properties. Using single-cell RNA sequencing, we comprehensively characterized the transcriptomes of most PN classes and unequivocally mapped transcriptomes to specific olfactory function for six classes. Transcriptomes of closely related PN classes exhibit the largest differences during circuit assembly but become indistinguishable in adults, suggesting that neuronal subtype diversity peaks during development. Transcription factors and cell-surface molecules are the most differentially expressed genes between classes and are highly informative in encoding cell identity, enabling us to identify a new lineage-specific transcription factor that instructs PN dendrite targeting. These findings establish that neuronal transcriptomic identity corresponds with anatomical and physiological identity defined by connectivity and function.
BACKGROUND: Among the most prominent molecular constituents of a recycling synaptic vesicle is the clathrin triskelion, composed of clathrin light chain (Clc) and clathrin heavy chain (Chc). Remarkably, it remains unknown whether clathrin is strictly necessary for the stimulus-dependent re-formation of a synaptic vesicle and, conversely, whether clathrin-independent vesicle endocytosis exists at the neuronal synapse. RESULTS: We employ FlAsH-FALI-mediated protein photoinactivation to rapidly (3 min) and specifically disrupt Clc function at the Drosophila neuromuscular junction. We first demonstrate that Clc photoinactivation does not impair synaptic-vesicle fusion. We then provide electrophysiological and ultrastructural evidence that synaptic vesicles, once fused with the plasma membrane, cannot be re-formed after Clc photoinactivation. Finally, we demonstrate that stimulus-dependent membrane internalization occurs after Clc photoinactivation. However, newly internalized membrane fails to resolve into synaptic vesicles. Rather, newly internalized membrane forms large and extensive internal-membrane compartments that are never observed at a wild-type synapse. CONCLUSIONS: We make three major conclusions. (1) FlAsH-FALI-mediated protein photoinactivation rapidly and specifically disrupts Clc function with no effect on synaptic-vesicle fusion. (2) Synaptic-vesicle re-formation does not occur after Clc photoinactivation. By extension, clathrin-independent "kiss-and-run" endocytosis does not sustain synaptic transmission during a stimulus train at this synapse. (3) Stimulus-dependent, clathrin-independent membrane internalization exists at this synapse, but it is unable to generate fusion-competent, small-diameter synaptic vesicles.
Preclinical animal models have provided strong evidence that estrogen (E) therapy (ET) enhances cognition and induces spinogenesis in neuronal circuits. However, clinical studies have been inconsistent, with some studies revealing adverse effects of ET, including an increased risk of dementia. In an effort to bridge this disconnect between the preclinical and clinical data, we have developed a nonhuman primate (NHP) model of ET combined with high-resolution dendritic spine analysis of dorsolateral prefrontal cortical (dlPFC) neurons. Previously, we reported cyclic ET in aged, ovariectomized NHPs increased spine density on dlPFC neurons. Here, we report that monkeys treated with cyclic E treatment paired with cyclic progesterone (P), continuous E combined with P (either cyclic or continuous), or unopposed continuous E failed to increase spines on dlPFC neurons. Given that the most prevalent form of ET prescribed to women is a combined and continuous E and P, these data bring into convergence the human neuropsychological findings and preclinical neurobiological evidence that standard hormone therapy in women is unlikely to yield the synaptic benefit presumed to underlie the cognitive enhancement reported in animal models.
Detecting meaningful structure in neural activity and connectivity data is challenging in the presence of hidden nonlinearities, where traditional eigenvalue-based methods may be misleading. We introduce a novel approach to matrix analysis, called clique topology, that extracts features of the data invariant under nonlinear monotone transformations. These features can be used to detect both random and geometric structure, and depend only on the relative ordering of matrix entries. We then analyzed the activity of pyramidal neurons in rat hippocampus, recorded while the animal was exploring a 2D environment, and confirmed that our method is able to detect geometric organization using only the intrinsic pattern of neural correlations. Remarkably, we found similar results during nonspatial behaviors such as wheel running and rapid eye movement (REM) sleep. This suggests that the geometric structure of correlations is shaped by the underlying hippocampal circuits and is not merely a consequence of position coding. We propose that clique topology is a powerful new tool for matrix analysis in biological settings, where the relationship of observed quantities to more meaningful variables is often nonlinear and unknown.
The antennal lobe (AL) is the primary structure in the Drosophila brain that relays odor information from the antennae to higher brain centers. The characterization of uniglomerular projection neurons (PNs) and some local interneurons has facilitated our understanding of olfaction; however, many other AL neurons remain unidentified. Because neuron types are mostly specified by lineage and temporal origins, we use the MARCM techniques with a set of enhancer-trap GAL4 lines to perform systematical lineage analysis to characterize neuron morphologies, lineage origin and birth timing in the three AL neuron lineages that contain GAL4-GH146-positive PNs: anterodorsal, lateral and ventral lineages. The results show that the anterodorsal lineage is composed of pure uniglomerular PNs that project through the inner antennocerebral tract. The ventral lineage produces uniglomerular and multiglomerular PNs that project through the middle antennocerebral tract. The lateral lineage generates multiple types of neurons, including uniglomeurlar PNs, diverse atypical PNs, various types of AL local interneurons and the neurons that make no connection within the ALs. Specific neuron types in all three lineages are produced in specific time windows, although multiple neuron types in the lateral lineage are made simultaneously. These systematic cell lineage analyses have not only filled gaps in the olfactory map, but have also exemplified additional strategies used in the brain to increase neuronal diversity.
BACKGROUND: The insect brain can be divided into neuropils that are formed by neurites of both local and remote origin. The complexity of the interconnections obscures how these neuropils are established and interconnected through development. The Drosophila central brain develops from a fixed number of neuroblasts (NBs) that deposit neurons in regional clusters. RESULTS: By determining individual NB clones and pursuing their projections into specific neuropils, we unravel the regional development of the brain neural network. Exhaustive clonal analysis revealed 95 stereotyped neuronal lineages with characteristic cell-body locations and neurite trajectories. Most clones show complex projection patterns, but despite the complexity, neighboring clones often coinnervate the same local neuropil or neuropils and further target a restricted set of distant neuropils. CONCLUSIONS: These observations argue for regional clonal development of both neuropils and neuropil connectivity throughout the Drosophila central brain.
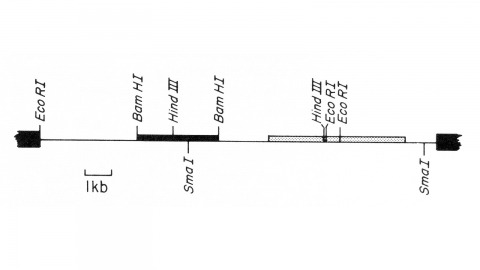
View Publication PageWe describe the isolation of a cloned DNA segment carrying unique sequences from the white locus of Drosophila melanogaster. Sequences within the cloned segment are shown to hybridize in situ to the white locus region on the polytene chromosomes of both wild-type strains and strains carrying chromosomal rearrangements whose breakpoints bracket the white locus. We further show that two small deficiency mutations, deleting white locus genetic elements but not those of complementation groups contiguous to white, delete the genomic sequences corresponding to a portion of the cloned segment. The strategy we have employed to isolate this cloned segment exploits the existence of an allele at the white locus containing a copy of a previously cloned transposable, reiterated DNA sequence element. We describe a simple, rapid method for retrieving cloned segments carrying a copy of the transposable element together with contiguous sequences corresponding to this allele. The strategy described is potentially general and we discuss its application to the cloning of the DNA sequences of other genes in Drosophila, including those identified only by genetic analysis and for which no RNA product is known.