Filter
Associated Lab
- Aguilera Castrejon Lab (17) Apply Aguilera Castrejon Lab filter
- Ahrens Lab (69) Apply Ahrens Lab filter
- Aso Lab (42) Apply Aso Lab filter
- Baker Lab (38) Apply Baker Lab filter
- Betzig Lab (115) Apply Betzig Lab filter
- Beyene Lab (14) Apply Beyene Lab filter
- Bock Lab (17) Apply Bock Lab filter
- Branson Lab (54) Apply Branson Lab filter
- Card Lab (43) Apply Card Lab filter
- Cardona Lab (64) Apply Cardona Lab filter
- Chklovskii Lab (13) Apply Chklovskii Lab filter
- Clapham Lab (15) Apply Clapham Lab filter
- Cui Lab (19) Apply Cui Lab filter
- Darshan Lab (12) Apply Darshan Lab filter
- Dennis Lab (2) Apply Dennis Lab filter
- Dickson Lab (46) Apply Dickson Lab filter
- Druckmann Lab (25) Apply Druckmann Lab filter
- Dudman Lab (53) Apply Dudman Lab filter
- Eddy/Rivas Lab (30) Apply Eddy/Rivas Lab filter
- Egnor Lab (11) Apply Egnor Lab filter
- Espinosa Medina Lab (21) Apply Espinosa Medina Lab filter
- Feliciano Lab (10) Apply Feliciano Lab filter
- Fetter Lab (41) Apply Fetter Lab filter
- FIB-SEM Technology (1) Apply FIB-SEM Technology filter
- Fitzgerald Lab (29) Apply Fitzgerald Lab filter
- Freeman Lab (15) Apply Freeman Lab filter
- Funke Lab (42) Apply Funke Lab filter
- Gonen Lab (91) Apply Gonen Lab filter
- Grigorieff Lab (62) Apply Grigorieff Lab filter
- Harris Lab (64) Apply Harris Lab filter
- Heberlein Lab (94) Apply Heberlein Lab filter
- Hermundstad Lab (30) Apply Hermundstad Lab filter
- Hess Lab (79) Apply Hess Lab filter
- Ilanges Lab (3) Apply Ilanges Lab filter
- Jayaraman Lab (48) Apply Jayaraman Lab filter
- Ji Lab (33) Apply Ji Lab filter
- Johnson Lab (6) Apply Johnson Lab filter
- Kainmueller Lab (19) Apply Kainmueller Lab filter
- Karpova Lab (14) Apply Karpova Lab filter
- Keleman Lab (13) Apply Keleman Lab filter
- Keller Lab (76) Apply Keller Lab filter
- Koay Lab (18) Apply Koay Lab filter
- Lavis Lab (154) Apply Lavis Lab filter
- Lee (Albert) Lab (34) Apply Lee (Albert) Lab filter
- Leonardo Lab (23) Apply Leonardo Lab filter
- Li Lab (30) Apply Li Lab filter
- Lippincott-Schwartz Lab (178) Apply Lippincott-Schwartz Lab filter
- Liu (Yin) Lab (7) Apply Liu (Yin) Lab filter
- Liu (Zhe) Lab (64) Apply Liu (Zhe) Lab filter
- Looger Lab (138) Apply Looger Lab filter
- Magee Lab (49) Apply Magee Lab filter
- Menon Lab (18) Apply Menon Lab filter
- Murphy Lab (13) Apply Murphy Lab filter
- O'Shea Lab (7) Apply O'Shea Lab filter
- Otopalik Lab (13) Apply Otopalik Lab filter
- Pachitariu Lab (49) Apply Pachitariu Lab filter
- Pastalkova Lab (18) Apply Pastalkova Lab filter
- Pavlopoulos Lab (19) Apply Pavlopoulos Lab filter
- Pedram Lab (15) Apply Pedram Lab filter
- Podgorski Lab (16) Apply Podgorski Lab filter
- Reiser Lab (54) Apply Reiser Lab filter
- Riddiford Lab (44) Apply Riddiford Lab filter
- Romani Lab (49) Apply Romani Lab filter
- Rubin Lab (148) Apply Rubin Lab filter
- Saalfeld Lab (64) Apply Saalfeld Lab filter
- Satou Lab (16) Apply Satou Lab filter
- Scheffer Lab (38) Apply Scheffer Lab filter
- Schreiter Lab (69) Apply Schreiter Lab filter
- Sgro Lab (21) Apply Sgro Lab filter
- Shroff Lab (31) Apply Shroff Lab filter
- Simpson Lab (23) Apply Simpson Lab filter
- Singer Lab (80) Apply Singer Lab filter
- Spruston Lab (97) Apply Spruston Lab filter
- Stern Lab (158) Apply Stern Lab filter
- Sternson Lab (54) Apply Sternson Lab filter
- Stringer Lab (39) Apply Stringer Lab filter
- Svoboda Lab (135) Apply Svoboda Lab filter
- Tebo Lab (35) Apply Tebo Lab filter
- Tervo Lab (9) Apply Tervo Lab filter
- Tillberg Lab (21) Apply Tillberg Lab filter
- Tjian Lab (64) Apply Tjian Lab filter
- Truman Lab (88) Apply Truman Lab filter
- Turaga Lab (53) Apply Turaga Lab filter
- Turner Lab (39) Apply Turner Lab filter
- Vale Lab (8) Apply Vale Lab filter
- Voigts Lab (4) Apply Voigts Lab filter
- Wang (Meng) Lab (27) Apply Wang (Meng) Lab filter
- Wang (Shaohe) Lab (25) Apply Wang (Shaohe) Lab filter
- Wu Lab (9) Apply Wu Lab filter
- Zlatic Lab (28) Apply Zlatic Lab filter
- Zuker Lab (25) Apply Zuker Lab filter
Associated Project Team
- CellMap (12) Apply CellMap filter
- COSEM (3) Apply COSEM filter
- FIB-SEM Technology (5) Apply FIB-SEM Technology filter
- Fly Descending Interneuron (12) Apply Fly Descending Interneuron filter
- Fly Functional Connectome (14) Apply Fly Functional Connectome filter
- Fly Olympiad (5) Apply Fly Olympiad filter
- FlyEM (56) Apply FlyEM filter
- FlyLight (50) Apply FlyLight filter
- GENIE (47) Apply GENIE filter
- Integrative Imaging (7) Apply Integrative Imaging filter
- Larval Olympiad (2) Apply Larval Olympiad filter
- MouseLight (18) Apply MouseLight filter
- NeuroSeq (1) Apply NeuroSeq filter
- ThalamoSeq (1) Apply ThalamoSeq filter
- Tool Translation Team (T3) (28) Apply Tool Translation Team (T3) filter
- Transcription Imaging (49) Apply Transcription Imaging filter
Publication Date
- 2025 (227) Apply 2025 filter
- 2024 (212) Apply 2024 filter
- 2023 (158) Apply 2023 filter
- 2022 (192) Apply 2022 filter
- 2021 (194) Apply 2021 filter
- 2020 (196) Apply 2020 filter
- 2019 (202) Apply 2019 filter
- 2018 (232) Apply 2018 filter
- 2017 (217) Apply 2017 filter
- 2016 (209) Apply 2016 filter
- 2015 (252) Apply 2015 filter
- 2014 (236) Apply 2014 filter
- 2013 (194) Apply 2013 filter
- 2012 (190) Apply 2012 filter
- 2011 (190) Apply 2011 filter
- 2010 (161) Apply 2010 filter
- 2009 (158) Apply 2009 filter
- 2008 (140) Apply 2008 filter
- 2007 (106) Apply 2007 filter
- 2006 (92) Apply 2006 filter
- 2005 (67) Apply 2005 filter
- 2004 (57) Apply 2004 filter
- 2003 (58) Apply 2003 filter
- 2002 (39) Apply 2002 filter
- 2001 (28) Apply 2001 filter
- 2000 (29) Apply 2000 filter
- 1999 (14) Apply 1999 filter
- 1998 (18) Apply 1998 filter
- 1997 (16) Apply 1997 filter
- 1996 (10) Apply 1996 filter
- 1995 (18) Apply 1995 filter
- 1994 (12) Apply 1994 filter
- 1993 (10) Apply 1993 filter
- 1992 (6) Apply 1992 filter
- 1991 (11) Apply 1991 filter
- 1990 (11) Apply 1990 filter
- 1989 (6) Apply 1989 filter
- 1988 (1) Apply 1988 filter
- 1987 (7) Apply 1987 filter
- 1986 (4) Apply 1986 filter
- 1985 (5) Apply 1985 filter
- 1984 (2) Apply 1984 filter
- 1983 (2) Apply 1983 filter
- 1982 (3) Apply 1982 filter
- 1981 (3) Apply 1981 filter
- 1980 (1) Apply 1980 filter
- 1979 (1) Apply 1979 filter
- 1976 (2) Apply 1976 filter
- 1973 (1) Apply 1973 filter
- 1970 (1) Apply 1970 filter
- 1967 (1) Apply 1967 filter
Type of Publication
4202 Publications
Showing 1531-1540 of 4202 resultsGap junctions (GJs) represent connexin-rich membrane domains that connect interiors of adjoining cells in mammalian tissues. How fast GJs can respond to bacterial pathogens has not been known previously. Using Bessel beam plane illumination and confocal spinning disk microscopy, we found fast (~500 ms) formation of connexin-depleted regions (CDRs) inside GJ plaques between cells exposed to AB5 toxins. CDR formation appears as a fast redistribution of connexin channels within GJ plaques with minor changes in outline or geometry. CDR formation does not depend on membrane trafficking or submembrane cytoskeleton and has no effect on GJ conductance. However, CDR responses depend on membrane lipids, can be modified by cholesterol-clustering agents and extracellular K(+) ion concentration, and influence cAMP signaling. The CDR response of GJ plaques to bacterial toxins is a phenomenon observed for all tested connexin isoforms. Through signaling, the CDR response may enable cells to sense exposure to AB5 toxins. CDR formation may reflect lipid-phase separation events in the biological membrane of the GJ plaque, leading to increased connexin packing and lipid reorganization. Our data demonstrate very fast dynamics (in the millisecond-to-second range) within GJ plaques, which previously were considered to be relatively stable, long-lived structures.
Cortical information processing is under state-dependent control of subcortical neuromodulatory systems. Although this modulatory effect is thought to be mediated mainly by slow nonsynaptic metabotropic receptors, other mechanisms, such as direct synaptic transmission, are possible. Yet, it is currently unknown if any such form of subcortical control exists. Here, we present direct evidence of a strong, spatiotemporally precise excitatory input from an ascending neuromodulatory center. Selective stimulation of serotonergic median raphe neurons produced a rapid activation of hippocampal interneurons. At the network level, this subcortical drive was manifested as a pattern of effective disynaptic GABAergic inhibition that spread throughout the circuit. This form of subcortical network regulation should be incorporated into current concepts of normal and pathological cortical function.
The comprehensive reconstruction of cell lineages in complex multicellular organisms is a central goal of developmental biology. We present an open-source computational framework for the segmentation and tracking of cell nuclei with high accuracy and speed. We demonstrate its (i) generality by reconstructing cell lineages in four-dimensional, terabyte-sized image data sets of fruit fly, zebrafish and mouse embryos acquired with three types of fluorescence microscopes, (ii) scalability by analyzing advanced stages of development with up to 20,000 cells per time point at 26,000 cells min(-1) on a single computer workstation and (iii) ease of use by adjusting only two parameters across all data sets and providing visualization and editing tools for efficient data curation. Our approach achieves on average 97.0% linkage accuracy across all species and imaging modalities. Using our system, we performed the first cell lineage reconstruction of early Drosophila melanogaster nervous system development, revealing neuroblast dynamics throughout an entire embryo.
Optical recording of intricate molecular dynamics is becoming an indispensable technique for biological studies, accelerated by the development of new or improved biosensors and microscopy technology. This creates major computational challenges to extract and quantify biologically meaningful spatiotemporal patterns embedded within complex and rich data sources, many of which cannot be captured with existing methods. Here, we introduce activity quantification and analysis (AQuA2), a fast, accurate, and versatile data analysis platform built upon advanced machine-learning techniques. It decomposes complex live-imaging-based datasets into elementary signaling events, allowing accurate and unbiased quantification of molecular activities and identification of consensus functional units. We demonstrate applications across a wide range of biosensors, cell types, organs, animal models, microscopy techniques, and imaging approaches. As exemplar findings, we show how AQuA2 identified drug-dependent interactions between neurons and astroglia, as well as distinct sensorimotor signal propagation patterns in the mouse spinal cord.
Recording light-microscopy images of large, nontransparent specimens, such as developing multicellular organisms, is complicated by decreased contrast resulting from light scattering. Early zebrafish development can be captured by standard light-sheet microscopy, but new imaging strategies are required to obtain high-quality data of late development or of less transparent organisms. We combined digital scanned laser light-sheet fluorescence microscopy with incoherent structured-illumination microscopy (DSLM-SI) and created structured-illumination patterns with continuously adjustable frequencies. Our method discriminates the specimen-related scattered background from signal fluorescence, thereby removing out-of-focus light and optimizing the contrast of in-focus structures. DSLM-SI provides rapid control of the illumination pattern, exceptional imaging quality, and high imaging speeds. We performed long-term imaging of zebrafish development for 58 h and fast multiple-view imaging of early Drosophila melanogaster development. We reconstructed cell positions over time from the Drosophila DSLM-SI data and created a fly digital embryo.
Understanding genetic and cellular bases of adult form remains a fundamental goal at the intersection of developmental and evolutionary biology. The skin pigment cells of vertebrates, derived from embryonic neural crest, are a useful system for elucidating mechanisms of fate specification, pattern formation, and how particular phenotypes impact organismal behavior and ecology. In a survey of fishes, including the zebrafish , we identified two populations of white pigment cells-leucophores-one of which arises by transdifferentiation of adult melanophores and another of which develops from a yellow-orange xanthophore or xanthophore-like progenitor. Single-cell transcriptomic, mutational, chemical, and ultrastructural analyses of zebrafish leucophores revealed cell-type-specific chemical compositions, organelle configurations, and genetic requirements. At the organismal level, we identified distinct physiological responses of leucophores during environmental background matching, and we showed that leucophore complement influences behavior. Together, our studies reveal independently arisen pigment cell types and mechanisms of fate acquisition in zebrafish and illustrate how concerted analyses across hierarchical levels can provide insights into phenotypes and their evolution.
Fatty acids (FAs) provide cellular energy under starvation, yet how they mobilize and move into mitochondria in starved cells, driving oxidative respiration, is unclear. Here, we clarify this process by visualizing FA trafficking with a fluorescent FA probe. The labeled FA accumulated in lipid droplets (LDs) in well-fed cells but moved from LDs into mitochondria when cells were starved. Autophagy in starved cells replenished LDs with FAs, increasing LD number over time. Cytoplasmic lipases removed FAs from LDs, enabling their transfer into mitochondria. This required mitochondria to be highly fused and localized near LDs. When mitochondrial fusion was prevented in starved cells, FAs neither homogeneously distributed within mitochondria nor became efficiently metabolized. Instead, FAs reassociated with LDs and fluxed into neighboring cells. Thus, FAs engage in complex trafficking itineraries regulated by cytoplasmic lipases, autophagy, and mitochondrial fusion dynamics, ensuring maximum oxidative metabolism and avoidance of FA toxicity in starved cells.
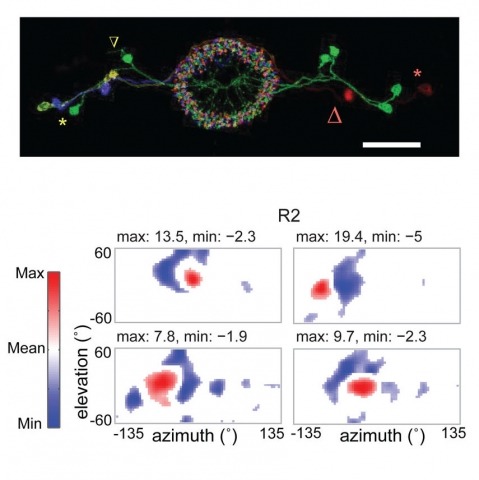
Many animals, including insects, are known to use visual landmarks to orient in their environment. In Drosophila melanogaster, behavioural genetics studies have identified a higher brain structure called the central complex as being required for the fly’s innate responses to vertical visual features and its short- and long-term memory for visual patterns. But whether and how neurons of the fly central complex represent visual features are unknown. Here we use two-photon calcium imaging in head-fixed walking and flying flies to probe visuomotor responses of ring neurons—a class of central complex neurons that have been implicated in landmark-driven spatial memory in walking flies and memory for visual patterns in tethered flying flies. We show that dendrites of ring neurons are visually responsive and arranged retinotopically. Ring neuron receptive fields comprise both excitatory and inhibitory subfields, resembling those of simple cells in the mammalian primary visual cortex. Ring neurons show strong and, in some cases, direction-selective orientation tuning, with a notable preference for vertically oriented features similar to those that evoke innate responses in flies. Visual responses were diminished during flight, but, in contrast with the hypothesized role of the central complex in the control of locomotion, not modulated during walking. Taken together, these results indicate that ring neurons represent behaviourally relevant visual features in the fly’s environment, enabling downstream central complex circuits to produce appropriate motor commands. More broadly, this study opens the door to mechanistic investigations of circuit computations underlying visually guided action selection in the Drosophila central complex.
Animals rely on dedicated sensory circuits to extract and encode environmental features. How individual neurons integrate and translate these features into behavioral responses remains a major question. Here, we identify a visual projection neuron type that conveys predator approach information to the Drosophila giant fiber (GF) escape circuit. Genetic removal of this input during looming stimuli reveals that it encodes angular expansion velocity, whereas other input cell type(s) encode angular size. Motor program selection and timing emerge from linear integration of these two features within the GF. Linear integration improves size detection invariance over prior models and appropriately biases motor selection to rapid, GF-mediated escapes during fast looms. Our findings suggest feature integration, and motor control may occur as simultaneous operations within the same neuron and establish the Drosophila escape circuit as a model system in which these computations may be further dissected at the circuit level.