Filter
Associated Lab
- Aguilera Castrejon Lab (19) Apply Aguilera Castrejon Lab filter
- Ahrens Lab (73) Apply Ahrens Lab filter
- Aso Lab (42) Apply Aso Lab filter
- Baker Lab (38) Apply Baker Lab filter
- Betzig Lab (116) Apply Betzig Lab filter
- Beyene Lab (15) Apply Beyene Lab filter
- Bock Lab (17) Apply Bock Lab filter
- Branson Lab (55) Apply Branson Lab filter
- Card Lab (43) Apply Card Lab filter
- Cardona Lab (64) Apply Cardona Lab filter
- Chklovskii Lab (13) Apply Chklovskii Lab filter
- Clapham Lab (16) Apply Clapham Lab filter
- Cui Lab (19) Apply Cui Lab filter
- Darshan Lab (12) Apply Darshan Lab filter
- Dennis Lab (3) Apply Dennis Lab filter
- Dickson Lab (46) Apply Dickson Lab filter
- Druckmann Lab (25) Apply Druckmann Lab filter
- Dudman Lab (56) Apply Dudman Lab filter
- Eddy/Rivas Lab (30) Apply Eddy/Rivas Lab filter
- Egnor Lab (11) Apply Egnor Lab filter
- Espinosa Medina Lab (23) Apply Espinosa Medina Lab filter
- Feliciano Lab (12) Apply Feliciano Lab filter
- Fetter Lab (41) Apply Fetter Lab filter
- FIB-SEM Technology (1) Apply FIB-SEM Technology filter
- Fitzgerald Lab (30) Apply Fitzgerald Lab filter
- Freeman Lab (15) Apply Freeman Lab filter
- Funke Lab (46) Apply Funke Lab filter
- Gonen Lab (91) Apply Gonen Lab filter
- Grigorieff Lab (62) Apply Grigorieff Lab filter
- Harris Lab (65) Apply Harris Lab filter
- Heberlein Lab (94) Apply Heberlein Lab filter
- Hermundstad Lab (30) Apply Hermundstad Lab filter
- Hess Lab (80) Apply Hess Lab filter
- Ilanges Lab (4) Apply Ilanges Lab filter
- Jayaraman Lab (48) Apply Jayaraman Lab filter
- Ji Lab (33) Apply Ji Lab filter
- Johnson Lab (6) Apply Johnson Lab filter
- Kainmueller Lab (19) Apply Kainmueller Lab filter
- Karpova Lab (15) Apply Karpova Lab filter
- Keleman Lab (13) Apply Keleman Lab filter
- Keller Lab (76) Apply Keller Lab filter
- Koay Lab (20) Apply Koay Lab filter
- Lavis Lab (158) Apply Lavis Lab filter
- Lee (Albert) Lab (34) Apply Lee (Albert) Lab filter
- Leonardo Lab (23) Apply Leonardo Lab filter
- Li Lab (32) Apply Li Lab filter
- Lippincott-Schwartz Lab (180) Apply Lippincott-Schwartz Lab filter
- Liu (Yin) Lab (8) Apply Liu (Yin) Lab filter
- Liu (Zhe) Lab (65) Apply Liu (Zhe) Lab filter
- Looger Lab (138) Apply Looger Lab filter
- Magee Lab (49) Apply Magee Lab filter
- Menon Lab (18) Apply Menon Lab filter
- Murphy Lab (13) Apply Murphy Lab filter
- O'Shea Lab (8) Apply O'Shea Lab filter
- Otopalik Lab (13) Apply Otopalik Lab filter
- Pachitariu Lab (54) Apply Pachitariu Lab filter
- Pastalkova Lab (19) Apply Pastalkova Lab filter
- Pavlopoulos Lab (19) Apply Pavlopoulos Lab filter
- Pedram Lab (15) Apply Pedram Lab filter
- Podgorski Lab (16) Apply Podgorski Lab filter
- Reiser Lab (55) Apply Reiser Lab filter
- Riddiford Lab (44) Apply Riddiford Lab filter
- Romani Lab (51) Apply Romani Lab filter
- Rubin Lab (149) Apply Rubin Lab filter
- Saalfeld Lab (64) Apply Saalfeld Lab filter
- Satou Lab (18) Apply Satou Lab filter
- Scheffer Lab (38) Apply Scheffer Lab filter
- Schreiter Lab (70) Apply Schreiter Lab filter
- Schulze Lab (1) Apply Schulze Lab filter
- Sgro Lab (23) Apply Sgro Lab filter
- Shroff Lab (31) Apply Shroff Lab filter
- Simpson Lab (23) Apply Simpson Lab filter
- Singer Lab (80) Apply Singer Lab filter
- Spruston Lab (98) Apply Spruston Lab filter
- Stern Lab (160) Apply Stern Lab filter
- Sternson Lab (54) Apply Sternson Lab filter
- Stringer Lab (43) Apply Stringer Lab filter
- Svoboda Lab (136) Apply Svoboda Lab filter
- Tebo Lab (35) Apply Tebo Lab filter
- Tervo Lab (10) Apply Tervo Lab filter
- Tillberg Lab (22) Apply Tillberg Lab filter
- Tjian Lab (64) Apply Tjian Lab filter
- Truman Lab (88) Apply Truman Lab filter
- Turaga Lab (53) Apply Turaga Lab filter
- Turner Lab (38) Apply Turner Lab filter
- Vale Lab (8) Apply Vale Lab filter
- Voigts Lab (4) Apply Voigts Lab filter
- Wang (Meng) Lab (29) Apply Wang (Meng) Lab filter
- Wang (Shaohe) Lab (25) Apply Wang (Shaohe) Lab filter
- Wong-Campos Lab (1) Apply Wong-Campos Lab filter
- Wu Lab (9) Apply Wu Lab filter
- Zlatic Lab (28) Apply Zlatic Lab filter
- Zuker Lab (25) Apply Zuker Lab filter
Associated Project Team
- CellMap (13) Apply CellMap filter
- COSEM (3) Apply COSEM filter
- FIB-SEM Technology (5) Apply FIB-SEM Technology filter
- Fly Descending Interneuron (12) Apply Fly Descending Interneuron filter
- Fly Functional Connectome (14) Apply Fly Functional Connectome filter
- Fly Olympiad (5) Apply Fly Olympiad filter
- FlyEM (56) Apply FlyEM filter
- FlyLight (50) Apply FlyLight filter
- GENIE (47) Apply GENIE filter
- Integrative Imaging (9) Apply Integrative Imaging filter
- Larval Olympiad (2) Apply Larval Olympiad filter
- MouseLight (18) Apply MouseLight filter
- NeuroSeq (1) Apply NeuroSeq filter
- ThalamoSeq (1) Apply ThalamoSeq filter
- Tool Translation Team (T3) (29) Apply Tool Translation Team (T3) filter
- Transcription Imaging (49) Apply Transcription Imaging filter
Publication Date
- 2026 (74) Apply 2026 filter
- 2025 (222) Apply 2025 filter
- 2024 (210) Apply 2024 filter
- 2023 (158) Apply 2023 filter
- 2022 (192) Apply 2022 filter
- 2021 (194) Apply 2021 filter
- 2020 (196) Apply 2020 filter
- 2019 (202) Apply 2019 filter
- 2018 (232) Apply 2018 filter
- 2017 (217) Apply 2017 filter
- 2016 (209) Apply 2016 filter
- 2015 (252) Apply 2015 filter
- 2014 (236) Apply 2014 filter
- 2013 (194) Apply 2013 filter
- 2012 (190) Apply 2012 filter
- 2011 (190) Apply 2011 filter
- 2010 (161) Apply 2010 filter
- 2009 (158) Apply 2009 filter
- 2008 (140) Apply 2008 filter
- 2007 (106) Apply 2007 filter
- 2006 (92) Apply 2006 filter
- 2005 (67) Apply 2005 filter
- 2004 (57) Apply 2004 filter
- 2003 (58) Apply 2003 filter
- 2002 (39) Apply 2002 filter
- 2001 (28) Apply 2001 filter
- 2000 (29) Apply 2000 filter
- 1999 (14) Apply 1999 filter
- 1998 (18) Apply 1998 filter
- 1997 (16) Apply 1997 filter
- 1996 (10) Apply 1996 filter
- 1995 (18) Apply 1995 filter
- 1994 (12) Apply 1994 filter
- 1993 (10) Apply 1993 filter
- 1992 (6) Apply 1992 filter
- 1991 (11) Apply 1991 filter
- 1990 (11) Apply 1990 filter
- 1989 (6) Apply 1989 filter
- 1988 (1) Apply 1988 filter
- 1987 (7) Apply 1987 filter
- 1986 (4) Apply 1986 filter
- 1985 (5) Apply 1985 filter
- 1984 (2) Apply 1984 filter
- 1983 (2) Apply 1983 filter
- 1982 (3) Apply 1982 filter
- 1981 (3) Apply 1981 filter
- 1980 (1) Apply 1980 filter
- 1979 (1) Apply 1979 filter
- 1976 (2) Apply 1976 filter
- 1973 (1) Apply 1973 filter
- 1970 (1) Apply 1970 filter
- 1967 (1) Apply 1967 filter
Type of Publication
4269 Publications
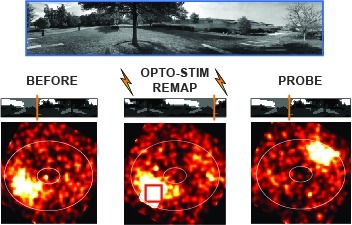
Showing 1681-1690 of 4269 resultsMany animals rely on an internal heading representation when navigating in varied environments. How this representation is linked to the sensory cues that define different surroundings is unclear. In the fly brain, heading is represented by 'compass' neurons that innervate a ring-shaped structure known as the ellipsoid body. Each compass neuron receives inputs from 'ring' neurons that are selective for particular visual features; this combination provides an ideal substrate for the extraction of directional information from a visual scene. Here we combine two-photon calcium imaging and optogenetics in tethered flying flies with circuit modelling, and show how the correlated activity of compass and visual neurons drives plasticity, which flexibly transforms two-dimensional visual cues into a stable heading representation. We also describe how this plasticity enables the fly to convert a partial heading representation, established from orienting within part of a novel setting, into a complete heading representation. Our results provide mechanistic insight into the memory-related computations that are essential for flexible navigation in varied surroundings.
Mammalian mitochondria maintain a small circular genome that encodes RNA and polypeptides that are essential for the generation of ATP through oxidative phosphorylation. The mechanism of replication of mammalian mitochondrial DNA (mtDNA) has recently been a topic of controversy. New evidence has led to a modified strand-displacement model that reconciles much of the current data. This revision stems from a new appreciation for alternative light-strand origins. We consider here some of the potential mechanisms for light-strand origin initiation. We also consider further the susceptibility of branch migration within replicating mtDNA molecules. The existence of alternative light-strand origins and a propensity for branch migration in replicating mtDNA molecules exposes a new array of possible configurations of mtDNA. The assortment and assignment of these forms is relevant to the interpretation of experimental data and may also yield insight into the molecular basis of replication errors.
Interindividual differences in neuronal wiring may contribute to behavioral individuality and affect susceptibility to neurological disorders. To investigate the causes and potential consequences of wiring variation in Drosophila melanogaster, we focused on a hemilineage of ventral nerve cord interneurons that exhibits morphological variability. We find that late-born subclasses of the 12A hemilineage are highly sensitive to genetic and environmental variation. Neurons in the second thoracic segment are particularly variable with regard to two developmental decisions, whereas its segmental homologs are more robust. This variability "hotspot" depends on Ultrabithorax expression in the 12A neurons, indicating variability is cell-intrinsic and under genetic control. 12A development is more variable and sensitive to temperature in long-established laboratory strains than in strains recently derived from the wild. Strains with a high frequency of one of the 12A variants also showed a high frequency of animals with delayed spontaneous flight initiation, whereas other wing-related behaviors did not show such a correlation and were thus not overtly affected by 12A variation. These results show that neurodevelopmental robustness is variable and under genetic control in Drosophila and suggest that the fly may serve as a model for identifying conserved gene pathways that stabilize wiring in stressful developmental environments. Moreover, some neuronal lineages are variation hotspots and thus may be more amenable to evolutionary change.
Genetically hard-wired neural mechanisms must enforce behavioral reproductive isolation because interspecies courtship is rare even in sexually na{\"ıve animals of most species. We find that the chemoreceptor Gr32a inhibits male D. melanogaster from courting diverse fruit fly species. Gr32a recognizes nonvolatile aversive cues present on these reproductively dead-end targets, and activity of Gr32a neurons is necessary and sufficient to inhibit interspecies courtship. Male-specific Fruitless (Fru(M)), a master regulator of courtship, also inhibits interspecies courtship. Gr32a and Fru(M) are not coexpressed, but Fru(M) neurons contact Gr32a neurons, suggesting that these genes influence a shared neural circuit that inhibits interspecies courtship. Gr32a and Fru(M) also suppress within-species intermale courtship, but we show that distinct mechanisms preclude sexual displays toward conspecific males and other species. Although this chemosensory pathway does not inhibit interspecies mating in D. melanogaster females, similar mechanisms appear to inhibit this behavior in many other male drosophilids.
Animals execute one particular behavior among many others in a context-dependent manner, yet the mechanisms underlying such behavioral choice remain poorly understood. Here we studied how two fundamental behaviors, sex and sleep, interact at genetic and neuronal levels in Drosophila. We show that an increased need for sleep inhibits male sexual behavior by decreasing the activity of the male-specific P1 neurons that coexpress the sex determination genes fru (M) and dsx, but does not affect female sexual behavior. Further, we delineate a sex-specific neuronal circuit wherein the P1 neurons encoding increased courtship drive suppressed male sleep by forming mutually excitatory connections with the fru (M) -positive sleep-controlling DN1 neurons. In addition, we find that FRU(M) regulates male courtship and sleep through distinct neural substrates. These studies reveal the genetic and neuronal basis underlying the sex-specific interaction between sleep and sexual behaviors in Drosophila, and provide insights into how competing behaviors are co-regulated.Genes and circuits involved in sleep and sexual arousal have been extensively studied in Drosophila. Here the authors identify the sex determination genes fruitless and doublesex, and a sex-specific P1-DN1 neuronal feedback that governs the interaction between these competing behaviors.
Species of the Drosophila melanogaster species subgroup, including the species D. simulans, D. mauritiana, D. yakuba, and D. santomea, have long served as model systems for studying evolution. Studies in these species have been limited, however, by a paucity of genetic and transgenic reagents. Here we describe a collection of transgenic and genetic strains generated to facilitate genetic studies within and between these species. We have generated many strains of each species containing mapped piggyBac transposons including an enhanced yellow fluorescent protein gene expressed in the eyes and a phiC31 attP site-specific integration site. We have tested a subset of these lines for integration efficiency and reporter gene expression levels. We have also generated a smaller collection of other lines expressing other genetically encoded fluorescent molecules in the eyes and a number of other transgenic reagents that will be useful for functional studies in these species. In addition, we have mapped the insertion locations of 58 transposable elements in D. virilis that will be useful for genetic mapping studies.
The transition from outcrossing to predominant self-fertilization is one of the most common evolutionary transitions in flowering plants. This shift is often accompanied by a suite of changes in floral and reproductive characters termed the selfing syndrome. Here, we characterize the genetic architecture and evolutionary forces underlying evolution of the selfing syndrome in Capsella rubella following its recent divergence from the outcrossing ancestor C. grandiflora. We conduct genotyping by multiplexed shotgun sequencing and map floral and reproductive traits in a large (N= 550) F2 population. Our results suggest that in contrast to previous studies of the selfing syndrome, changes at a few loci, some with major effects, have shaped the evolution of the selfing syndrome in Capsella. The directionality of QTL effects, as well as population genetic patterns of polymorphism and divergence at 318 loci, is consistent with a history of directional selection on the selfing syndrome. Our study is an important step toward characterizing the genetic basis and evolutionary forces underlying the evolution of the selfing syndrome in a genetically accessible model system.
Male sexual characters are often among the first traits to diverge between closely related species and identifying the genetic basis of such changes can contribute to our understanding of their evolutionary history. However, little is known about the genetic architecture or the specific genes underlying the evolution of male genitalia. The morphology of the claspers, posterior lobes and anal plates exhibit striking differences between Drosophila mauritiana and Drosophila simulans. Using QTL and introgression-based high-resolution mapping, we identified several small regions on chromosome arms 3L and 3R that contribute to differences in these traits. However, we found that the loci underlying the evolution of clasper differences between these two species are independent from those that contribute to posterior lobe and anal plate divergence. Furthermore, while most of the loci affect each trait in the same direction and act additively, we also found evidence for epistasis between loci for clasper bristle number. In addition, we conducted an RNAi screen in D. melanogaster to investigate if positional and expression candidate genes located on chromosome 3L, are also involved in genital development. We found that six of these genes, including components of Wnt signaling and male-specific lethal 3 (msl3), regulate the development of genital traits consistent with the effects of the introgressed regions where they are located and that thus represent promising candidate genes for the evolution these traits.
Considerable progress has been made over the past couple of decades concerning the molecular bases of neurobehavioral function and dysfunction. The field of neurobehavioral genetics is becoming mature. Genetic factors contributing to neurologic diseases such as Alzheimer's disease have been found and evidence for genetic factors contributing to other diseases such as schizophrenia and autism are likely. This genetic approach can also benefit the field of behavioral neurotoxicology. It is clear that there is substantial heterogeneity of response with behavioral impairments resulting from neurotoxicants. Many factors contribute to differential sensitivity, but it is likely that genetic variability plays a prominent role. Important discoveries concerning genetics and behavioral neurotoxicity are being made on a broad front from work with invertebrate and piscine mutant models to classic mouse knockout models and human epidemiologic studies of polymorphisms. Discovering genetic factors of susceptibility to neurobehavioral toxicity not only helps identify those at special risk, it also advances our understanding of the mechanisms by which toxicants impair neurobehavioral function in the larger population. This symposium organized by Edward Levin and Annette Kirshner, brought together researchers from the laboratories of Michael Aschner, Douglas Ruden, Ulrike Heberlein, Edward Levin and Kathleen Welsh-Bohmer conducting studies with Caenorhabditis elegans, Drosophila, fish, rodents and humans studies to determine the role of genetic factors in susceptibility to behavioral impairment from neurotoxic exposure.
BACKGROUND: In most organisms in which acute ethanol exposure has been studied, it leads to similar changes in behavior. Generally, low ethanol doses activate the central nervous system, whereas high doses are sedative. Sensitivity to the acute intoxicating effects of ethanol is in part under genetic control in rodents and humans, and reduced sensitivity in humans predicts the development of alcoholism (Crabbe et al., 1994; Schuckit, 1994). We have established Drosophila melanogaster as a model organism to study the mechanisms that regulate acute sensitivity to ethanol. METHODS: We measured the effects of ethanol vapor on Drosophila locomotor behaviors by using three different assays. Horizontal locomotion was quantified in a locomotor chamber, turning behavior was assayed in narrow tubes, and ethanol-induced loss of postural control was measured in an inebriometer. Mutants with altered sensitivity to the acute effects of ethanol were generated by treatment with ethyl methane sulfonate and isolated by selection in the inebriometer. We ascertained the effects of these mutations on ethanol pharmacokinetics by measuring ethanol levels in extracts of flies at various times during and after ethanol exposure. RESULTS: Among nearly 30,000 potentially mutant flies tested, we isolated 19 mutant strains with reduced and 4 strains with increased sensitivity to the acute effects of ethanol as measured in the inebriometer. Of these mutants, four showed changes in ethanol absorption. Two mutants, named barfly and tipsy to reflect their reduced and increased ethanol sensitivity in the inebriometer, respectively, were analyzed for locomotor behaviors. Both mutants exhibited ethanol-induced hyperactivity that was indistinguishable from wild type. However, barfly and tipsy displayed reduced and increased sensitivity to the sedative effects of ethanol, respectively. Finally, both mutants showed an increased rate of ethanol-induced turning behavior. CONCLUSIONS: The effects of acute ethanol exposure on Drosophila locomotor behaviors are remarkably similar to those described for mammals. The analysis of mutants with altered sensitivity to ethanol revealed that the genetic pathways which regulate these responses are complex and that single genes can affect hyperactivity, turning, and sedation independently.