Filter
Associated Lab
- Aguilera Castrejon Lab (17) Apply Aguilera Castrejon Lab filter
- Ahrens Lab (69) Apply Ahrens Lab filter
- Aso Lab (42) Apply Aso Lab filter
- Baker Lab (38) Apply Baker Lab filter
- Betzig Lab (115) Apply Betzig Lab filter
- Beyene Lab (14) Apply Beyene Lab filter
- Bock Lab (17) Apply Bock Lab filter
- Branson Lab (54) Apply Branson Lab filter
- Card Lab (43) Apply Card Lab filter
- Cardona Lab (64) Apply Cardona Lab filter
- Chklovskii Lab (13) Apply Chklovskii Lab filter
- Clapham Lab (15) Apply Clapham Lab filter
- Cui Lab (19) Apply Cui Lab filter
- Darshan Lab (12) Apply Darshan Lab filter
- Dennis Lab (2) Apply Dennis Lab filter
- Dickson Lab (46) Apply Dickson Lab filter
- Druckmann Lab (25) Apply Druckmann Lab filter
- Dudman Lab (53) Apply Dudman Lab filter
- Eddy/Rivas Lab (30) Apply Eddy/Rivas Lab filter
- Egnor Lab (11) Apply Egnor Lab filter
- Espinosa Medina Lab (21) Apply Espinosa Medina Lab filter
- Feliciano Lab (10) Apply Feliciano Lab filter
- Fetter Lab (41) Apply Fetter Lab filter
- FIB-SEM Technology (1) Apply FIB-SEM Technology filter
- Fitzgerald Lab (29) Apply Fitzgerald Lab filter
- Freeman Lab (15) Apply Freeman Lab filter
- Funke Lab (42) Apply Funke Lab filter
- Gonen Lab (91) Apply Gonen Lab filter
- Grigorieff Lab (62) Apply Grigorieff Lab filter
- Harris Lab (64) Apply Harris Lab filter
- Heberlein Lab (94) Apply Heberlein Lab filter
- Hermundstad Lab (30) Apply Hermundstad Lab filter
- Hess Lab (79) Apply Hess Lab filter
- Ilanges Lab (3) Apply Ilanges Lab filter
- Jayaraman Lab (48) Apply Jayaraman Lab filter
- Ji Lab (33) Apply Ji Lab filter
- Johnson Lab (6) Apply Johnson Lab filter
- Kainmueller Lab (19) Apply Kainmueller Lab filter
- Karpova Lab (14) Apply Karpova Lab filter
- Keleman Lab (13) Apply Keleman Lab filter
- Keller Lab (76) Apply Keller Lab filter
- Koay Lab (18) Apply Koay Lab filter
- Lavis Lab (154) Apply Lavis Lab filter
- Lee (Albert) Lab (34) Apply Lee (Albert) Lab filter
- Leonardo Lab (23) Apply Leonardo Lab filter
- Li Lab (30) Apply Li Lab filter
- Lippincott-Schwartz Lab (178) Apply Lippincott-Schwartz Lab filter
- Liu (Yin) Lab (7) Apply Liu (Yin) Lab filter
- Liu (Zhe) Lab (64) Apply Liu (Zhe) Lab filter
- Looger Lab (138) Apply Looger Lab filter
- Magee Lab (49) Apply Magee Lab filter
- Menon Lab (18) Apply Menon Lab filter
- Murphy Lab (13) Apply Murphy Lab filter
- O'Shea Lab (7) Apply O'Shea Lab filter
- Otopalik Lab (13) Apply Otopalik Lab filter
- Pachitariu Lab (49) Apply Pachitariu Lab filter
- Pastalkova Lab (18) Apply Pastalkova Lab filter
- Pavlopoulos Lab (19) Apply Pavlopoulos Lab filter
- Pedram Lab (15) Apply Pedram Lab filter
- Podgorski Lab (16) Apply Podgorski Lab filter
- Reiser Lab (54) Apply Reiser Lab filter
- Riddiford Lab (44) Apply Riddiford Lab filter
- Romani Lab (49) Apply Romani Lab filter
- Rubin Lab (148) Apply Rubin Lab filter
- Saalfeld Lab (64) Apply Saalfeld Lab filter
- Satou Lab (16) Apply Satou Lab filter
- Scheffer Lab (38) Apply Scheffer Lab filter
- Schreiter Lab (69) Apply Schreiter Lab filter
- Sgro Lab (21) Apply Sgro Lab filter
- Shroff Lab (31) Apply Shroff Lab filter
- Simpson Lab (23) Apply Simpson Lab filter
- Singer Lab (80) Apply Singer Lab filter
- Spruston Lab (97) Apply Spruston Lab filter
- Stern Lab (158) Apply Stern Lab filter
- Sternson Lab (54) Apply Sternson Lab filter
- Stringer Lab (39) Apply Stringer Lab filter
- Svoboda Lab (135) Apply Svoboda Lab filter
- Tebo Lab (35) Apply Tebo Lab filter
- Tervo Lab (9) Apply Tervo Lab filter
- Tillberg Lab (21) Apply Tillberg Lab filter
- Tjian Lab (64) Apply Tjian Lab filter
- Truman Lab (88) Apply Truman Lab filter
- Turaga Lab (53) Apply Turaga Lab filter
- Turner Lab (39) Apply Turner Lab filter
- Vale Lab (8) Apply Vale Lab filter
- Voigts Lab (4) Apply Voigts Lab filter
- Wang (Meng) Lab (27) Apply Wang (Meng) Lab filter
- Wang (Shaohe) Lab (25) Apply Wang (Shaohe) Lab filter
- Wu Lab (9) Apply Wu Lab filter
- Zlatic Lab (28) Apply Zlatic Lab filter
- Zuker Lab (25) Apply Zuker Lab filter
Associated Project Team
- CellMap (12) Apply CellMap filter
- COSEM (3) Apply COSEM filter
- FIB-SEM Technology (5) Apply FIB-SEM Technology filter
- Fly Descending Interneuron (12) Apply Fly Descending Interneuron filter
- Fly Functional Connectome (14) Apply Fly Functional Connectome filter
- Fly Olympiad (5) Apply Fly Olympiad filter
- FlyEM (56) Apply FlyEM filter
- FlyLight (50) Apply FlyLight filter
- GENIE (47) Apply GENIE filter
- Integrative Imaging (7) Apply Integrative Imaging filter
- Larval Olympiad (2) Apply Larval Olympiad filter
- MouseLight (18) Apply MouseLight filter
- NeuroSeq (1) Apply NeuroSeq filter
- ThalamoSeq (1) Apply ThalamoSeq filter
- Tool Translation Team (T3) (28) Apply Tool Translation Team (T3) filter
- Transcription Imaging (49) Apply Transcription Imaging filter
Publication Date
- 2025 (227) Apply 2025 filter
- 2024 (212) Apply 2024 filter
- 2023 (158) Apply 2023 filter
- 2022 (192) Apply 2022 filter
- 2021 (194) Apply 2021 filter
- 2020 (196) Apply 2020 filter
- 2019 (202) Apply 2019 filter
- 2018 (232) Apply 2018 filter
- 2017 (217) Apply 2017 filter
- 2016 (209) Apply 2016 filter
- 2015 (252) Apply 2015 filter
- 2014 (236) Apply 2014 filter
- 2013 (194) Apply 2013 filter
- 2012 (190) Apply 2012 filter
- 2011 (190) Apply 2011 filter
- 2010 (161) Apply 2010 filter
- 2009 (158) Apply 2009 filter
- 2008 (140) Apply 2008 filter
- 2007 (106) Apply 2007 filter
- 2006 (92) Apply 2006 filter
- 2005 (67) Apply 2005 filter
- 2004 (57) Apply 2004 filter
- 2003 (58) Apply 2003 filter
- 2002 (39) Apply 2002 filter
- 2001 (28) Apply 2001 filter
- 2000 (29) Apply 2000 filter
- 1999 (14) Apply 1999 filter
- 1998 (18) Apply 1998 filter
- 1997 (16) Apply 1997 filter
- 1996 (10) Apply 1996 filter
- 1995 (18) Apply 1995 filter
- 1994 (12) Apply 1994 filter
- 1993 (10) Apply 1993 filter
- 1992 (6) Apply 1992 filter
- 1991 (11) Apply 1991 filter
- 1990 (11) Apply 1990 filter
- 1989 (6) Apply 1989 filter
- 1988 (1) Apply 1988 filter
- 1987 (7) Apply 1987 filter
- 1986 (4) Apply 1986 filter
- 1985 (5) Apply 1985 filter
- 1984 (2) Apply 1984 filter
- 1983 (2) Apply 1983 filter
- 1982 (3) Apply 1982 filter
- 1981 (3) Apply 1981 filter
- 1980 (1) Apply 1980 filter
- 1979 (1) Apply 1979 filter
- 1976 (2) Apply 1976 filter
- 1973 (1) Apply 1973 filter
- 1970 (1) Apply 1970 filter
- 1967 (1) Apply 1967 filter
Type of Publication
4202 Publications
Showing 421-430 of 4202 resultsOrganismal aging involves functional declines in both somatic and reproductive tissues. Multiple strategies have been discovered to extend lifespan across species. However, how age-related molecular changes differ among various tissues and how those lifespan-extending strategies slow tissue aging in distinct manners remain unclear. Here we generated the transcriptomic Cell Atlas of Worm Aging (CAWA, http://mengwanglab.org/atlas ) of wild-type and long-lived strains. We discovered cell-specific, age-related molecular and functional signatures across all somatic and germ cell types. We developed transcriptomic aging clocks for different tissues and quantitatively determined how three different pro-longevity strategies slow tissue aging distinctively. Furthermore, through genome-wide profiling of alternative polyadenylation (APA) events in different tissues, we discovered cell-type-specific APA changes during aging and revealed how these changes are differentially affected by the pro-longevity strategies. Together, this study offers fundamental molecular insights into both somatic and reproductive aging and provides a valuable resource for in-depth understanding of the diversity of pro-longevity mechanisms.
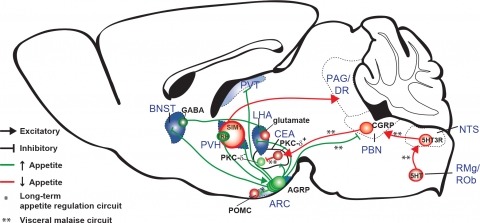
New tools for mapping and manipulating molecularly defined neural circuits have improved understanding of how the central nervous system regulates appetite. Studies focused on AGRP neurons, a starvation-sensitive hypothalamic population, have identified multiple circuit elements that can elicit or suppress feeding behavior. Distinct axon projections of this neuron population point to different circuits that regulate long-term appetite, short-term feeding, or visceral malaise-mediated anorexia. Here, we review recent studies examining these neural circuits that control food intake. © 2014 S. Karger AG, Basel.
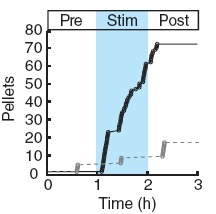
Two intermingled hypothalamic neuron populations specified by expression of agouti-related peptide (AGRP) or pro-opiomelanocortin (POMC) positively and negatively influence feeding behavior, respectively, possibly by reciprocally regulating downstream melanocortin receptors. However, the sufficiency of these neurons to control behavior and the relationship of their activity to the magnitude and dynamics of feeding are unknown. To measure this, we used channelrhodopsin-2 for cell type-specific photostimulation. Activation of only 800 AGRP neurons in mice evoked voracious feeding within minutes. The behavioral response increased with photoexcitable neuron number, photostimulation frequency and stimulus duration. Conversely, POMC neuron stimulation reduced food intake and body weight, which required melanocortin receptor signaling. However, AGRP neuron-mediated feeding was not dependent on suppressing this melanocortin pathway, indicating that AGRP neurons directly engage feeding circuits. Furthermore, feeding was evoked selectively over drinking without training or prior photostimulus exposure, which suggests that AGRP neurons serve a dedicated role coordinating this complex behavior.
A decline in ocular lens transparency known as cataract afflicts 90% of individuals by the age 70. Chronic deterioration of lens tissue occurs as a pathophysiological consequence of defective water and nutrient circulation through channel and transporter proteins. A key component is the aquaporin-0 (AQP0) water channel whose permeability is tightly regulated in healthy lenses. Using a variety of cellular and biochemical approaches we have discovered that products of the A-kinase anchoring protein 2 gene (AKAP2/AKAP-KL) form a stable complex with AQP0 to sequester protein kinase A (PKA) with the channel. This permits PKA phosphorylation of serine 235 within a calmodulin (CaM)-binding domain of AQP0. The additional negative charge introduced by phosphoserine 235 perturbs electrostatic interactions between AQP0 and CaM to favour water influx through the channel. In isolated mouse lenses, displacement of PKA from the AKAP2-AQP0 channel complex promotes cortical cataracts as characterized by severe opacities and cellular damage. Thus, anchored PKA modulation of AQP0 is a homeostatic mechanism that must be physically intact to preserve lens transparency.
Camera networks are widely used for tasks such as surveillance, monitoring and tracking. In order to accomplish these tasks, knowledge of localization information such as camera locations and other geometric constraints about the environment (e.g. walls, rooms, and building layout) are typically considered to be essential. However, this information is not always required for many tasks such as estimating the topology of camera network coverage, or coordinate-free object tracking and navigation. In this paper, we propose a simplicial representation (called CN- complex) that can be constructed from discrete local observations from cameras, and utilize this novel representation to recover the topological information of the network coverage. We prove that our representation captures the correct topological information from network coverage for 2.5-D layouts, and demonstrate their utility in simulations as well as a real-world experimental set-up. Our proposed approach is particularly useful in the context of ad-hoc camera networks in indoor/outdoor urban environments with distributed but limited computational power and energy.
To thrive, organisms must maintain physiological and environmental variables in suitable ranges. Given that these variables undergo constant fluctuations over varying time scales, how do biological control systems maintain control over these values? We explored this question in the context of phototactic behavior in larval zebrafish. We demonstrate that larval zebrafish use phototaxis to maintain environmental luminance at a set point, that the value of this set point fluctuates on a time scale of seconds when environmental luminance changes, and that it is determined by calculating the mean input across both sides of the visual field. These results expand on previous studies of flexible phototaxis in larval zebrafish; they suggest that larval zebrafish exert homeostatic control over the luminance of their surroundings, and that feedback from the surroundings drives allostatic changes to the luminance set point. As such, we describe a novel behavioral algorithm with which larval zebrafish exert control over a sensory variable.
Data acquisition of cryo-electron tomography (CET) samples described in previous chapters involves relatively imprecise mechanical motions: the tilt series has shifts, rotations, and several other distortions between projections. Alignment is the procedure of correcting for these effects in each image and requires the estimation of a projection model that describes how points from the sample in three-dimensions are projected to generate two-dimensional images. This estimation is enabled by finding corresponding common features between images. This chapter reviews several software packages that perform alignment and reconstruction tasks completely automatically (or with minimal user intervention) in two main scenarios: using gold fiducial markers as high contrast features or using relevant biological structures present in the image (marker-free). In particular, we emphasize the key decision points in the process that users should focus on in order to obtain high-resolution reconstructions.
3D reconstruction from serial 2D microscopy images depends on non-linear alignment of serial sections. For some structures, such as the neuronal circuitry of the brain, very large images at very high resolution are necessary to permit reconstruction. These very large images prevent the direct use of classical registration methods. We propose in this work a method to deal with the non-linear alignment of arbitrarily large 2D images using the finite support properties of cubic B-splines. After initial affine alignment, each large image is split into a grid of smaller overlapping sub-images, which are individually registered using cubic B-splines transformations. Inside the overlapping regions between neighboring sub-images, the coefficients of the knots controlling the B-splines deformations are blended, to create a virtual large grid of knots for the whole image. The sub-images are resampled individually, using the new coefficients, and assembled together into a final large aligned image. We evaluated the method on a series of large transmission electron microscopy images and our results indicate significant improvements compared to both manual and affine alignment.
Previously, we showed that the mouse LIM-domain only 4 (Lmo4) gene, which encodes a protein containing two zinc-finger LIM domains that interact with various DNA-binding transcription factors, attenuates behavioral sensitivity to repeated cocaine administration. Here we show that transcription of anaplastic lymphoma kinase (Alk) is repressed by LMO4 in the striatum and that Alk promotes the development of cocaine sensitization and conditioned place preference, a measure of cocaine reward. Since LMO4 is known to interact with estrogen receptor α (ERα) at the promoters of target genes, we investigated whether Alk expression might be controlled by a similar mechanism. We found that LMO4 and ERα are associated with the Alk promoter by chromatin immunoprecipitation and that Alk is an estrogen-responsive gene in the striatum. Moreover, we show that ERα knock-out mice exhibit enhanced cocaine sensitization and conditioned place preference and an increase in Alk expression in the nucleus accumbens. These data define a novel regulatory network involved in behavioral responses to cocaine. Interestingly, sex differences in several behavioral responses to cocaine in humans and rodents have been described, and estrogen is thought to mediate some of these differences. Our data suggest that estrogen regulation of Alk may be one mechanism responsible for sexually dimorphic responses to cocaine.