Filter
Associated Lab
- Aguilera Castrejon Lab (4) Apply Aguilera Castrejon Lab filter
- Ahrens Lab (63) Apply Ahrens Lab filter
- Aso Lab (42) Apply Aso Lab filter
- Baker Lab (19) Apply Baker Lab filter
- Betzig Lab (104) Apply Betzig Lab filter
- Beyene Lab (10) Apply Beyene Lab filter
- Bock Lab (14) Apply Bock Lab filter
- Branson Lab (52) Apply Branson Lab filter
- Card Lab (37) Apply Card Lab filter
- Cardona Lab (45) Apply Cardona Lab filter
- Chklovskii Lab (10) Apply Chklovskii Lab filter
- Clapham Lab (15) Apply Clapham Lab filter
- Cui Lab (19) Apply Cui Lab filter
- Darshan Lab (8) Apply Darshan Lab filter
- Dennis Lab (2) Apply Dennis Lab filter
- Dickson Lab (32) Apply Dickson Lab filter
- Druckmann Lab (21) Apply Druckmann Lab filter
- Dudman Lab (46) Apply Dudman Lab filter
- Eddy/Rivas Lab (30) Apply Eddy/Rivas Lab filter
- Egnor Lab (4) Apply Egnor Lab filter
- Espinosa Medina Lab (20) Apply Espinosa Medina Lab filter
- Feliciano Lab (14) Apply Feliciano Lab filter
- Fetter Lab (31) Apply Fetter Lab filter
- FIB-SEM Technology (1) Apply FIB-SEM Technology filter
- Fitzgerald Lab (17) Apply Fitzgerald Lab filter
- Freeman Lab (15) Apply Freeman Lab filter
- Funke Lab (46) Apply Funke Lab filter
- Gonen Lab (59) Apply Gonen Lab filter
- Grigorieff Lab (34) Apply Grigorieff Lab filter
- Harris Lab (55) Apply Harris Lab filter
- Heberlein Lab (13) Apply Heberlein Lab filter
- Hermundstad Lab (28) Apply Hermundstad Lab filter
- Hess Lab (77) Apply Hess Lab filter
- Ilanges Lab (4) Apply Ilanges Lab filter
- Jayaraman Lab (45) Apply Jayaraman Lab filter
- Ji Lab (33) Apply Ji Lab filter
- Johnson Lab (2) Apply Johnson Lab filter
- Karpova Lab (14) Apply Karpova Lab filter
- Keleman Lab (8) Apply Keleman Lab filter
- Keller Lab (61) Apply Keller Lab filter
- Koay Lab (4) Apply Koay Lab filter
- Lavis Lab (147) Apply Lavis Lab filter
- Lee (Albert) Lab (29) Apply Lee (Albert) Lab filter
- Leonardo Lab (19) Apply Leonardo Lab filter
- Li Lab (8) Apply Li Lab filter
- Lippincott-Schwartz Lab (108) Apply Lippincott-Schwartz Lab filter
- Liu (Yin) Lab (3) Apply Liu (Yin) Lab filter
- Liu (Zhe) Lab (60) Apply Liu (Zhe) Lab filter
- Looger Lab (137) Apply Looger Lab filter
- Magee Lab (31) Apply Magee Lab filter
- Menon Lab (12) Apply Menon Lab filter
- Murphy Lab (6) Apply Murphy Lab filter
- O'Shea Lab (7) Apply O'Shea Lab filter
- Otopalik Lab (1) Apply Otopalik Lab filter
- Pachitariu Lab (42) Apply Pachitariu Lab filter
- Pastalkova Lab (6) Apply Pastalkova Lab filter
- Pavlopoulos Lab (7) Apply Pavlopoulos Lab filter
- Pedram Lab (4) Apply Pedram Lab filter
- Podgorski Lab (16) Apply Podgorski Lab filter
- Reiser Lab (49) Apply Reiser Lab filter
- Riddiford Lab (20) Apply Riddiford Lab filter
- Romani Lab (40) Apply Romani Lab filter
- Rubin Lab (111) Apply Rubin Lab filter
- Saalfeld Lab (48) Apply Saalfeld Lab filter
- Satou Lab (3) Apply Satou Lab filter
- Scheffer Lab (38) Apply Scheffer Lab filter
- Schreiter Lab (53) Apply Schreiter Lab filter
- Schulze Lab (1) Apply Schulze Lab filter
- Sgro Lab (3) Apply Sgro Lab filter
- Shroff Lab (33) Apply Shroff Lab filter
- Simpson Lab (18) Apply Simpson Lab filter
- Singer Lab (37) Apply Singer Lab filter
- Spruston Lab (62) Apply Spruston Lab filter
- Stern Lab (77) Apply Stern Lab filter
- Sternson Lab (47) Apply Sternson Lab filter
- Stringer Lab (40) Apply Stringer Lab filter
- Svoboda Lab (132) Apply Svoboda Lab filter
- Tebo Lab (12) Apply Tebo Lab filter
- Tervo Lab (10) Apply Tervo Lab filter
- Tillberg Lab (19) Apply Tillberg Lab filter
- Tjian Lab (17) Apply Tjian Lab filter
- Truman Lab (58) Apply Truman Lab filter
- Turaga Lab (41) Apply Turaga Lab filter
- Turner Lab (27) Apply Turner Lab filter
- Vale Lab (8) Apply Vale Lab filter
- Voigts Lab (5) Apply Voigts Lab filter
- Wang (Meng) Lab (30) Apply Wang (Meng) Lab filter
- Wang (Shaohe) Lab (6) Apply Wang (Shaohe) Lab filter
- Wong-Campos Lab (1) Apply Wong-Campos Lab filter
- Wu Lab (8) Apply Wu Lab filter
- Zlatic Lab (26) Apply Zlatic Lab filter
- Zuker Lab (5) Apply Zuker Lab filter
Associated Project Team
- CellMap (13) Apply CellMap filter
- COSEM (3) Apply COSEM filter
- FIB-SEM Technology (5) Apply FIB-SEM Technology filter
- Fly Descending Interneuron (12) Apply Fly Descending Interneuron filter
- Fly Functional Connectome (14) Apply Fly Functional Connectome filter
- Fly Olympiad (5) Apply Fly Olympiad filter
- FlyEM (56) Apply FlyEM filter
- FlyLight (50) Apply FlyLight filter
- GENIE (47) Apply GENIE filter
- Integrative Imaging (10) Apply Integrative Imaging filter
- Larval Olympiad (2) Apply Larval Olympiad filter
- MouseLight (18) Apply MouseLight filter
- NeuroSeq (1) Apply NeuroSeq filter
- ThalamoSeq (1) Apply ThalamoSeq filter
- Tool Translation Team (T3) (29) Apply Tool Translation Team (T3) filter
- Transcription Imaging (45) Apply Transcription Imaging filter
Associated Support Team
- Project Pipeline Support (5) Apply Project Pipeline Support filter
- Anatomy and Histology (18) Apply Anatomy and Histology filter
- Cryo-Electron Microscopy (46) Apply Cryo-Electron Microscopy filter
- Electron Microscopy (18) Apply Electron Microscopy filter
- Gene Targeting and Transgenics (11) Apply Gene Targeting and Transgenics filter
- High Performance Computing (7) Apply High Performance Computing filter
- Integrative Imaging (24) Apply Integrative Imaging filter
- Invertebrate Shared Resource (40) Apply Invertebrate Shared Resource filter
- Janelia Experimental Technology (40) Apply Janelia Experimental Technology filter
- Management Team (1) Apply Management Team filter
- Mass Spectrometry (1) Apply Mass Spectrometry filter
- Molecular Genomics (15) Apply Molecular Genomics filter
- Project Technical Resources (54) Apply Project Technical Resources filter
- Quantitative Genomics (20) Apply Quantitative Genomics filter
- Scientific Computing (104) Apply Scientific Computing filter
- Stem Cell & Primary Culture (14) Apply Stem Cell & Primary Culture filter
- Viral Tools (14) Apply Viral Tools filter
- Vivarium (7) Apply Vivarium filter
Publication Date
- 2026 (95) Apply 2026 filter
- 2025 (221) Apply 2025 filter
- 2024 (209) Apply 2024 filter
- 2023 (157) Apply 2023 filter
- 2022 (166) Apply 2022 filter
- 2021 (175) Apply 2021 filter
- 2020 (177) Apply 2020 filter
- 2019 (177) Apply 2019 filter
- 2018 (206) Apply 2018 filter
- 2017 (186) Apply 2017 filter
- 2016 (191) Apply 2016 filter
- 2015 (195) Apply 2015 filter
- 2014 (190) Apply 2014 filter
- 2013 (136) Apply 2013 filter
- 2012 (112) Apply 2012 filter
- 2011 (98) Apply 2011 filter
- 2010 (61) Apply 2010 filter
- 2009 (56) Apply 2009 filter
- 2008 (40) Apply 2008 filter
- 2007 (21) Apply 2007 filter
- 2006 (3) Apply 2006 filter
2872 Janelia Publications
Showing 2151-2160 of 2872 resultsRecordings of the physiological history of cells provide insights into biological processes, yet obtaining such recordings is a challenge. To address this, we introduce a method to record transient cellular events for later analysis. We designed proteins that become labeled in the presence of both a specific cellular activity and a fluorescent substrate. The recording period is set by the presence of the substrate, whereas the cellular activity controls the degree of the labeling. The use of distinguishable substrates enabled the recording of successive periods of activity. We recorded protein-protein interactions, G protein-coupled receptor activation, and increases in intracellular calcium. Recordings of elevated calcium levels allowed selections of cells from heterogeneous populations for transcriptomic analysis and tracking of neuronal activities in flies and zebrafish.
Insects have evolved sophisticated reflexes to right themselves in mid-air. Their recovery mechanisms involve complex interactions among the physical senses, muscles, body, and wings, and they must obey the laws of flight. We sought to understand the key mechanisms involved in dragonfly righting reflexes and to develop physics-based models for understanding the control strategies of flight maneuvers. Using kinematic analyses, physical modeling, and three-dimensional flight simulations, we found that a dragonfly uses left-right wing pitch asymmetry to roll its body 180 degrees to recover from falling upside down in ~200 milliseconds. Experiments of dragonflies with blocked vision further revealed that this rolling maneuver is initiated by their ocelli and compound eyes. These results suggest a pathway from the dragonfly's visual system to the muscles regulating wing pitch that underly the recovery. The methods developed here offer quantitative tools for inferring insects' internal actions from their acrobatics, and are applicable to a broad class of natural and robotic flying systems.
Maintaining proper tension is critical for the organization and function of the plasma membrane. To study the mechanisms by which yeast restores normal plasma membrane tension, we used a microfluidics device to expose yeast to hyperosmotic conditions, which reduced cell volume and caused a ∼20% drop in cell surface area. The resulting low tension plasma membrane exhibited large clusters of negatively-charged glycerophospholipids together with nutrient transporters, suggesting phase segregation of the membrane. We found that endocytosis was blocked by the phase segregation and thus was not involved in removing excess membrane. In contrast, rapid recovery of plasma membrane tension was dependent on 1) eisosome morphology changes that were able to absorb most of the excess surface area and 2) lipid transport from the plasma membrane to the endoplasmic reticulum, where lipids were shunted into newly formed lipid droplets.
Neural computation involves diverse types of GABAergic inhibitory interneurons that are integrated with excitatory (E) neurons into precisely structured circuits. To understand how each neuron type shapes sensory representations, we measured firing patterns of defined types of neurons in the barrel cortex while mice performed an active, whisker-dependent object localization task. Touch excited fast-spiking (FS) interneurons at short latency, followed by activation of E neurons and somatostatin-expressing (SST) interneurons. Touch only weakly modulated vasoactive intestinal polypeptide-expressing (VIP) interneurons. Voluntary whisker movement activated FS neurons in the ventral posteromedial nucleus (VPM) target layers, a subset of SST neurons and a majority of VIP neurons. Together, FS neurons track thalamic input, mediating feedforward inhibition. SST neurons monitor local excitation, providing feedback inhibition. VIP neurons are activated by non-sensory inputs, disinhibiting E and FS neurons. Our data reveal rules of recruitment for interneuron types during behavior, providing foundations for understanding computation in cortical microcircuits.
Dopaminergic neurons (DANs) drive learning across the animal kingdom, but the upstream circuits that regulate their activity and thereby learning remain poorly understood. We provide a synaptic-resolution connectome of the circuitry upstream of all DANs in a learning center, the mushroom body of Drosophila larva. We discover afferent sensory pathways and a large population of neurons that provide feedback from mushroom body output neurons and link distinct memory systems (aversive and appetitive). We combine this with functional studies of DANs and their presynaptic partners and with comprehensive circuit modeling. We find that DANs compare convergent feedback from aversive and appetitive systems, which enables the computation of integrated predictions that may improve future learning. Computational modeling reveals that the discovered feedback motifs increase model flexibility and performance on learning tasks. Our study provides the most detailed view to date of biological circuit motifs that support associative learning.
Most cortical synapses are local and excitatory. Local recurrent circuits could implement amplification, allowing pattern completion and other computations. Cortical circuits contain subnetworks that consist of neurons with similar receptive fields and increased connectivity relative to the network average. Cortical neurons that encode different types of information are spatially intermingled and distributed over large brain volumes, and this complexity has hindered attempts to probe the function of these subnetworks by perturbing them individually. Here we use computational modelling, optical recordings and manipulations to probe the function of recurrent coupling in layer 2/3 of the mouse vibrissal somatosensory cortex during active tactile discrimination. A neural circuit model of layer 2/3 revealed that recurrent excitation enhances sensory signals by amplification, but only for subnetworks with increased connectivity. Model networks with high amplification were sensitive to damage: loss of a few members of the subnetwork degraded stimulus encoding. We tested this prediction by mapping neuronal selectivity and photoablating neurons with specific selectivity. Ablation of a small proportion of layer 2/3 neurons (10-20, less than 5% of the total) representing touch markedly reduced responses in the spared touch representation, but not in other representations. Ablations most strongly affected neurons with stimulus responses that were similar to those of the ablated population, which is also consistent with network models. Recurrence among cortical neurons with similar selectivity therefore drives input-specific amplification during behaviour.
The first step to probing any potential interaction between two biomolecules is to determine their spatial association. In other words, if two biomolecules localize similarly within a cell, then it is plausible they could interact. Traditionally, this is quantified through various colocalization metrics. These measures infer this association by estimating the degree to which fluorescent signals from each biomolecule overlap or correlate. However, these metrics are, at best, proxies, and they depend strongly on various experimental choices. Here, we define a new strategy that leverages multispectral imaging and phasor analysis, termed the phasor mixing coefficient (PMC). The PMC measures the precise mixing of fluorescent signals in each pixel. We demonstrate how the PMC captures complex biological subtlety by offering two distinct values, a global measure of overall color mixing and the homogeneity thereof. We additionally show that the PMC exhibits less sensitivity to signal-to-noise ratio, intensity threshold and background signal compared to canonical methods. Moreover, this method provides a means to visualize color mixing at each pixel. We show that the PMC offers users a nuanced and robust metric to quantify biological association.
The first step to probing any potential interaction between two biomolecules is to determine their spatial association. In other words, if two biomolecules localize similarly within a cell, then it is plausible they could interact. Traditionally, this is quantified through various colocalization metrics. These measures infer this association by estimating the degree to which fluorescent signals from each biomolecule overlap or correlate. However, these metrics are, at best, proxies, and they depend strongly on various experimental choices. Alternatively, here we define a new strategy which leverages multispectral imaging and phasor analysis, termed the Phasor Mixing Coefficient (PMC). PMC measures the precise mixing of fluorescent signals in each pixel. We demonstrate how PMC captures complex biological subtlety by offering two distinct values, a global measure of overall color mixing and the homogeneity thereof. We additionally show that PMC exhibits less sensitivity to signal-to-noise ratio, intensity threshold, and background signal compared to canonical methods. Moreover, this method provides a means to visualize color mixing at each pixel. We show that PMC offers users a nuanced and robust metric to quantify biological association.
Endocytic recycling of synaptic vesicles after exocytosis is critical for nervous system function. At synapses of cultured neurons that lack the two "neuronal" dynamins, dynamin 1 and 3, smaller excitatory postsynaptic currents are observed due to an impairment of the fission reaction of endocytosis that results in an accumulation of arrested clathrin-coated pits and a greatly reduced synaptic vesicle number. Surprisingly, despite a smaller readily releasable vesicle pool and fewer docked vesicles, a strong facilitation, which correlated with lower vesicle release probability, was observed upon action potential stimulation at such synapses. Furthermore, although network activity in mutant cultures was lower, Ca(2+)/calmodulin-dependent protein kinase II (CaMKII) activity was unexpectedly increased, consistent with the previous report of an enhanced state of synapsin 1 phosphorylation at CaMKII-dependent sites in such neurons. These changes were partially reversed by overnight silencing of synaptic activity with tetrodotoxin, a treatment that allows progression of arrested endocytic pits to synaptic vesicles. Facilitation was also counteracted by CaMKII inhibition. These findings reveal a mechanism aimed at preventing synaptic transmission failure due to vesicle depletion when recycling vesicle traffic is backed up by a defect in dynamin-dependent endocytosis and provide new insight into the coupling between endocytosis and exocytosis.
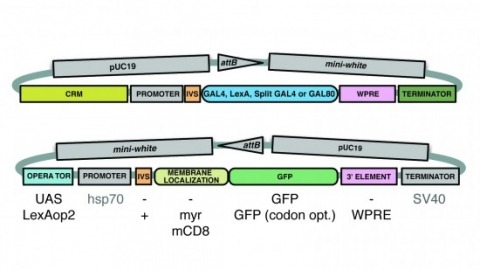
A wide variety of biological experiments rely on the ability to express an exogenous gene in a transgenic animal at a defined level and in a spatially and temporally controlled pattern. We describe major improvements of the methods available for achieving this objective in Drosophila melanogaster. We have systematically varied core promoters, UTRs, operator sequences, and transcriptional activating domains used to direct gene expression with the GAL4, LexA, and Split GAL4 transcription factors and the GAL80 transcriptional repressor. The use of site-specific integration allowed us to make quantitative comparisons between different constructs inserted at the same genomic location. We also characterized a set of PhiC31 integration sites for their ability to support transgene expression of both drivers and responders in the nervous system. The increased strength and reliability of these optimized reagents overcome many of the previous limitations of these methods and will facilitate genetic manipulations of greater complexity and sophistication.