Filter
Associated Lab
- Aguilera Castrejon Lab (17) Apply Aguilera Castrejon Lab filter
- Ahrens Lab (68) Apply Ahrens Lab filter
- Aso Lab (42) Apply Aso Lab filter
- Baker Lab (38) Apply Baker Lab filter
- Betzig Lab (115) Apply Betzig Lab filter
- Beyene Lab (14) Apply Beyene Lab filter
- Bock Lab (17) Apply Bock Lab filter
- Branson Lab (54) Apply Branson Lab filter
- Card Lab (43) Apply Card Lab filter
- Cardona Lab (64) Apply Cardona Lab filter
- Chklovskii Lab (13) Apply Chklovskii Lab filter
- Clapham Lab (15) Apply Clapham Lab filter
- Cui Lab (19) Apply Cui Lab filter
- Darshan Lab (12) Apply Darshan Lab filter
- Dennis Lab (1) Apply Dennis Lab filter
- Dickson Lab (46) Apply Dickson Lab filter
- Druckmann Lab (25) Apply Druckmann Lab filter
- Dudman Lab (52) Apply Dudman Lab filter
- Eddy/Rivas Lab (30) Apply Eddy/Rivas Lab filter
- Egnor Lab (11) Apply Egnor Lab filter
- Espinosa Medina Lab (20) Apply Espinosa Medina Lab filter
- Feliciano Lab (8) Apply Feliciano Lab filter
- Fetter Lab (41) Apply Fetter Lab filter
- FIB-SEM Technology (1) Apply FIB-SEM Technology filter
- Fitzgerald Lab (29) Apply Fitzgerald Lab filter
- Freeman Lab (15) Apply Freeman Lab filter
- Funke Lab (41) Apply Funke Lab filter
- Gonen Lab (91) Apply Gonen Lab filter
- Grigorieff Lab (62) Apply Grigorieff Lab filter
- Harris Lab (64) Apply Harris Lab filter
- Heberlein Lab (94) Apply Heberlein Lab filter
- Hermundstad Lab (29) Apply Hermundstad Lab filter
- Hess Lab (79) Apply Hess Lab filter
- Ilanges Lab (2) Apply Ilanges Lab filter
- Jayaraman Lab (47) Apply Jayaraman Lab filter
- Ji Lab (33) Apply Ji Lab filter
- Johnson Lab (6) Apply Johnson Lab filter
- Kainmueller Lab (19) Apply Kainmueller Lab filter
- Karpova Lab (14) Apply Karpova Lab filter
- Keleman Lab (13) Apply Keleman Lab filter
- Keller Lab (76) Apply Keller Lab filter
- Koay Lab (18) Apply Koay Lab filter
- Lavis Lab (152) Apply Lavis Lab filter
- Lee (Albert) Lab (34) Apply Lee (Albert) Lab filter
- Leonardo Lab (23) Apply Leonardo Lab filter
- Li Lab (29) Apply Li Lab filter
- Lippincott-Schwartz Lab (174) Apply Lippincott-Schwartz Lab filter
- Liu (Yin) Lab (7) Apply Liu (Yin) Lab filter
- Liu (Zhe) Lab (64) Apply Liu (Zhe) Lab filter
- Looger Lab (138) Apply Looger Lab filter
- Magee Lab (49) Apply Magee Lab filter
- Menon Lab (18) Apply Menon Lab filter
- Murphy Lab (13) Apply Murphy Lab filter
- O'Shea Lab (7) Apply O'Shea Lab filter
- Otopalik Lab (13) Apply Otopalik Lab filter
- Pachitariu Lab (49) Apply Pachitariu Lab filter
- Pastalkova Lab (18) Apply Pastalkova Lab filter
- Pavlopoulos Lab (19) Apply Pavlopoulos Lab filter
- Pedram Lab (15) Apply Pedram Lab filter
- Podgorski Lab (16) Apply Podgorski Lab filter
- Reiser Lab (52) Apply Reiser Lab filter
- Riddiford Lab (44) Apply Riddiford Lab filter
- Romani Lab (48) Apply Romani Lab filter
- Rubin Lab (146) Apply Rubin Lab filter
- Saalfeld Lab (64) Apply Saalfeld Lab filter
- Satou Lab (16) Apply Satou Lab filter
- Scheffer Lab (38) Apply Scheffer Lab filter
- Schreiter Lab (68) Apply Schreiter Lab filter
- Sgro Lab (21) Apply Sgro Lab filter
- Shroff Lab (31) Apply Shroff Lab filter
- Simpson Lab (23) Apply Simpson Lab filter
- Singer Lab (80) Apply Singer Lab filter
- Spruston Lab (94) Apply Spruston Lab filter
- Stern Lab (158) Apply Stern Lab filter
- Sternson Lab (54) Apply Sternson Lab filter
- Stringer Lab (39) Apply Stringer Lab filter
- Svoboda Lab (135) Apply Svoboda Lab filter
- Tebo Lab (34) Apply Tebo Lab filter
- Tervo Lab (9) Apply Tervo Lab filter
- Tillberg Lab (21) Apply Tillberg Lab filter
- Tjian Lab (64) Apply Tjian Lab filter
- Truman Lab (88) Apply Truman Lab filter
- Turaga Lab (52) Apply Turaga Lab filter
- Turner Lab (39) Apply Turner Lab filter
- Vale Lab (8) Apply Vale Lab filter
- Voigts Lab (3) Apply Voigts Lab filter
- Wang (Meng) Lab (23) Apply Wang (Meng) Lab filter
- Wang (Shaohe) Lab (25) Apply Wang (Shaohe) Lab filter
- Wu Lab (9) Apply Wu Lab filter
- Zlatic Lab (28) Apply Zlatic Lab filter
- Zuker Lab (25) Apply Zuker Lab filter
Associated Project Team
- CellMap (12) Apply CellMap filter
- COSEM (3) Apply COSEM filter
- FIB-SEM Technology (5) Apply FIB-SEM Technology filter
- Fly Descending Interneuron (12) Apply Fly Descending Interneuron filter
- Fly Functional Connectome (14) Apply Fly Functional Connectome filter
- Fly Olympiad (5) Apply Fly Olympiad filter
- FlyEM (56) Apply FlyEM filter
- FlyLight (50) Apply FlyLight filter
- GENIE (47) Apply GENIE filter
- Integrative Imaging (6) Apply Integrative Imaging filter
- Larval Olympiad (2) Apply Larval Olympiad filter
- MouseLight (18) Apply MouseLight filter
- NeuroSeq (1) Apply NeuroSeq filter
- ThalamoSeq (1) Apply ThalamoSeq filter
- Tool Translation Team (T3) (27) Apply Tool Translation Team (T3) filter
- Transcription Imaging (49) Apply Transcription Imaging filter
Publication Date
- 2025 (196) Apply 2025 filter
- 2024 (212) Apply 2024 filter
- 2023 (159) Apply 2023 filter
- 2022 (192) Apply 2022 filter
- 2021 (194) Apply 2021 filter
- 2020 (196) Apply 2020 filter
- 2019 (202) Apply 2019 filter
- 2018 (232) Apply 2018 filter
- 2017 (217) Apply 2017 filter
- 2016 (209) Apply 2016 filter
- 2015 (252) Apply 2015 filter
- 2014 (236) Apply 2014 filter
- 2013 (194) Apply 2013 filter
- 2012 (190) Apply 2012 filter
- 2011 (190) Apply 2011 filter
- 2010 (161) Apply 2010 filter
- 2009 (158) Apply 2009 filter
- 2008 (140) Apply 2008 filter
- 2007 (106) Apply 2007 filter
- 2006 (92) Apply 2006 filter
- 2005 (67) Apply 2005 filter
- 2004 (57) Apply 2004 filter
- 2003 (58) Apply 2003 filter
- 2002 (39) Apply 2002 filter
- 2001 (28) Apply 2001 filter
- 2000 (29) Apply 2000 filter
- 1999 (14) Apply 1999 filter
- 1998 (18) Apply 1998 filter
- 1997 (16) Apply 1997 filter
- 1996 (10) Apply 1996 filter
- 1995 (18) Apply 1995 filter
- 1994 (12) Apply 1994 filter
- 1993 (10) Apply 1993 filter
- 1992 (6) Apply 1992 filter
- 1991 (11) Apply 1991 filter
- 1990 (11) Apply 1990 filter
- 1989 (6) Apply 1989 filter
- 1988 (1) Apply 1988 filter
- 1987 (7) Apply 1987 filter
- 1986 (4) Apply 1986 filter
- 1985 (5) Apply 1985 filter
- 1984 (2) Apply 1984 filter
- 1983 (2) Apply 1983 filter
- 1982 (3) Apply 1982 filter
- 1981 (3) Apply 1981 filter
- 1980 (1) Apply 1980 filter
- 1979 (1) Apply 1979 filter
- 1976 (2) Apply 1976 filter
- 1973 (1) Apply 1973 filter
- 1970 (1) Apply 1970 filter
- 1967 (1) Apply 1967 filter
Type of Publication
4172 Publications
Showing 1961-1970 of 4172 resultsAlthough most proteins conform to the classical one-structure/one-function paradigm, an increasing number of proteins with dual structures and functions have been discovered. In response to cellular stimuli, such proteins undergo structural changes sufficiently dramatic to remodel even their secondary structures and domain organization. This "fold-switching" capability fosters protein multi-functionality, enabling cells to establish tight control over various biochemical processes. Accurate predictions of fold-switching proteins could both suggest underlying mechanisms for uncharacterized biological processes and reveal potential drug targets. Recently, we developed a prediction method for fold-switching proteins using structure-based thermodynamic calculations and discrepancies between predicted and experimentally determined protein secondary structure. Here we seek to leverage the negative information found in these secondary structure prediction discrepancies. To do this, we quantified secondary structure prediction accuracies of 192 known fold-switching regions (FSRs) within solved protein structures found in the Protein Data Bank (PDB). We find that the secondary structure prediction accuracies for these FSRs vary widely. Inaccurate secondary structure predictions are strongly associated with fold-switching proteins compared to equally long segments of non-fold-switching proteins selected at random. These inaccurate predictions are enriched in helix-to-strand and strand-to-coil discrepancies. Finally, we find that most proteins with inaccurate secondary structure predictions are underrepresented in the PDB compared with their alternatively folded cognates, suggesting that unequal representation of fold-switching conformers within the PDB could be an important cause of inaccurate secondary structure predictions. These results demonstrate that inconsistent secondary structure predictions can serve as a useful preliminary marker of fold switching. This article is protected by copyright. All rights reserved.
The final cleavage event that terminates cell division, abscission of the small, dense intercellular bridge, has been particularly challenging to resolve. Here, we describe imaging innovations that helped answer long-standing questions about the mechanism of abscission. We further explain how computational modeling of high-resolution data was employed to test hypotheses and generate additional insights. We present the model that emerges from application of these complimentary approaches. Similar experimental strategies will undoubtedly reveal exciting details about other underresolved cellular structures.
Dynamic modulation of the actin cytoskeleton is critical for synaptic plasticity, abnormalities of which are thought to contribute to mental illness and addiction. Here we report that mice lacking Eps8, a regulator of actin dynamics, are resistant to some acute intoxicating effects of ethanol and show increased ethanol consumption. In the brain, the N-methyl-D-aspartate (NMDA) receptor is a major target of ethanol. We show that Eps8 is localized to postsynaptic structures and is part of the NMDA receptor complex. Moreover, in Eps8 null mice, NMDA receptor currents and their sensitivity to inhibition by ethanol are abnormal. In addition, Eps8 null neurons are resistant to the actin-remodeling activities of NMDA and ethanol. We propose that proper regulation of the actin cytoskeleton is a key determinant of cellular and behavioral responses to ethanol.
Freshly isolated, depolarized rat hepatocytes can repolarize into bile canalicular networks when plated in collagen sandwich cultures. We studied the events underlying this repolarization process, focusing on how hepatocytes restore ATP synthesis and resupply biosynthetic precursors after the stress of being isolated from liver. We found that soon after being plated in collagen sandwich cultures, hepatocytes converted their mitochondria into highly fused networks. This occurred through a combination of upregulation of mitochondrial fusion proteins and downregulation of a mitochondrial fission protein. Mitochondria also became more active for oxidative phosphorylation, leading to overall increased ATP levels within cells. We further observed that autophagy was upregulated in the repolarizing hepatocytes. Boosted autophagy levels likely served to recycle cellular precursors, supplying building blocks for repolarization. Repolarizing hepatocytes also extensively degraded lipid droplets, whose fatty acids provide precursors for ?-oxidation to fuel oxidative phosphorylation in mitochondria. Thus, through coordination of mitochondrial fusion, autophagy, and lipid droplet consumption, depolarized hepatocytes are able to boost ATP synthesis and biosynthetic precursors to efficiently repolarize in collagen sandwich cultures.
The endoplasmic reticulum (ER) is an expansive, membrane-enclosed organelle that plays crucial roles in numerous cellular functions. We used emerging superresolution imaging technologies to clarify the morphology and dynamics of the peripheral ER, which contacts and modulates most other intracellular organelles. Peripheral components of the ER have classically been described as comprising both tubules and flat sheets. We show that this system consists almost exclusively of tubules at varying densities, including structures that we term ER matrices. Conventional optical imaging technologies had led to misidentification of these structures as sheets because of the dense clustering of tubular junctions and a previously uncharacterized rapid form of ER motion. The existence of ER matrices explains previous confounding evidence that had indicated the occurrence of ER “sheet” proliferation after overexpression of tubular junction–forming proteins.
In Drosophila photoreceptors the multivalent PDZ protein INAD organizes the phototransduction cascade into a macromolecular signaling complex containing the effector PLC, the light-activated TRP channels, and a regulatory PKC. Previously, we showed that the subcellular localization of INAD signaling complexes is critical for signaling. Now we have examined how INAD complexes are anchored and assembled in photoreceptor cells. We find that trp mutants, or transgenic flies expressing inaD alleles that disrupt the interaction between INAD and TRP, cause the mislocalization of the entire transduction complex. The INAD-TRP interaction is not required for targeting but rather for anchoring of complexes, because INAD and TRP can be targeted independently of each other. We also show that, in addition to its scaffold role, INAD functions to preassemble transduction complexes. Preassembly of signaling complexes helps to ensure that transduction complexes with the appropriate composition end up in the proper location. This may be a general mechanism used by cells to target different signaling machinery to the pertinent subcellular location.
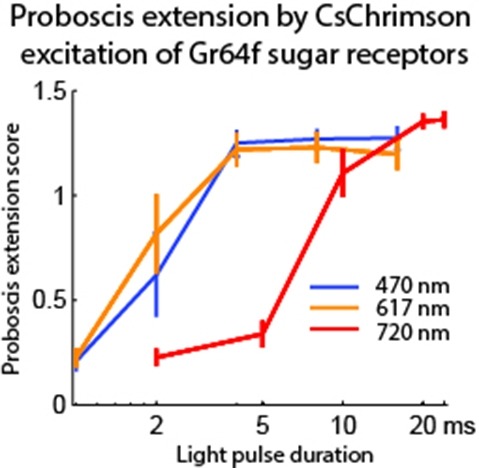
Optogenetic tools enable examination of how specific cell types contribute to brain circuit functions. A long-standing question is whether it is possible to independently activate two distinct neural populations in mammalian brain tissue. Such a capability would enable the study of how different synapses or pathways interact to encode information in the brain. Here we describe two channelrhodopsins, Chronos and Chrimson, discovered through sequencing and physiological characterization of opsins from over 100 species of alga. Chrimson’s excitation spectrum is red shifted by 45 nm relative to previous channelrhodopsins and can enable experiments in which red light is preferred. We show minimal visual system-mediated behavioral interference when using Chrimson in neurobehavioral studies in Drosophila melanogaster. Chronos has faster kinetics than previous channelrhodopsins yet is effectively more light sensitive. Together these two reagents enable two-color activation of neural spiking and downstream synaptic transmission in independent neural populations without detectable cross-talk in mouse brain slice.
Responses to threat-related stimuli are influenced by conscious and unconscious processes, but the neural systems underlying these processes and their relationship to anxiety have not been clearly delineated. Using fMRI, we investigated the neural responses associated with the conscious and unconscious (backwardly masked) perception of fearful faces in healthy volunteers who varied in threat sensitivity (Spielberger trait anxiety scale). Unconscious processing modulated activity only in the basolateral subregion of the amygdala, while conscious processing modulated activity only in the dorsal amygdala (containing the central nucleus). Whereas activation of the dorsal amygdala by conscious stimuli was consistent across subjects and independent of trait anxiety, activity in the basolateral amygdala to unconscious stimuli, and subjects’ reaction times, were predicted by individual differences in trait anxiety. These findings provide a biological basis for the unconscious emotional vigilance characteristic of anxiety and a means for investigating the mechanisms and efficacy of treatments for anxiety.
Polymorphisms in the inducible nitric oxide synthase gene (NOS2) promoter have been associated with clinical outcome from malaria. These include a CCTTT repeat (CCTTTn) 2.5 kilobases upstream from the NOS2 transcription start site, and two single nucleotide substitutions: G–>C at position -954 (G-954C), and C–>T at position -1173 (C-1173T). Although hypothesized to influence NO production in vivo, the functional relevance of (CCTTT)n and G-954C is uncertain because disease association studies have yielded inconsistent results. This study found no association between CCTTT repeat number and levels of plasma NO metabolites or peripheral blood mononuclear cell NOS activity in a cohort of asymptomatic malaria-exposed coastal Papua New Guineans 1-60 years old. This suggests that (CCTTT)n does not independently influence NOS2 transcription in vivo. Neither the G-954C nor the C-1173T polymorphisms were identified in this population, indicating the variability and complexity of selection for NOS2 promoter polymorphisms in different malaria-endemic populations.