Filter
Associated Lab
- Aso Lab (30) Apply Aso Lab filter
- Betzig Lab (1) Apply Betzig Lab filter
- Bock Lab (2) Apply Bock Lab filter
- Branson Lab (9) Apply Branson Lab filter
- Card Lab (5) Apply Card Lab filter
- Clapham Lab (1) Apply Clapham Lab filter
- Dickson Lab (2) Apply Dickson Lab filter
- Druckmann Lab (1) Apply Druckmann Lab filter
- Fetter Lab (1) Apply Fetter Lab filter
- Funke Lab (2) Apply Funke Lab filter
- Harris Lab (3) Apply Harris Lab filter
- Heberlein Lab (2) Apply Heberlein Lab filter
- Hermundstad Lab (2) Apply Hermundstad Lab filter
- Hess Lab (6) Apply Hess Lab filter
- Jayaraman Lab (7) Apply Jayaraman Lab filter
- Lippincott-Schwartz Lab (1) Apply Lippincott-Schwartz Lab filter
- Looger Lab (2) Apply Looger Lab filter
- O'Shea Lab (1) Apply O'Shea Lab filter
- Otopalik Lab (1) Apply Otopalik Lab filter
- Reiser Lab (17) Apply Reiser Lab filter
- Riddiford Lab (1) Apply Riddiford Lab filter
- Romani Lab (2) Apply Romani Lab filter
- Rubin Lab (149) Apply Rubin Lab filter
- Saalfeld Lab (5) Apply Saalfeld Lab filter
- Scheffer Lab (8) Apply Scheffer Lab filter
- Schreiter Lab (1) Apply Schreiter Lab filter
- Simpson Lab (3) Apply Simpson Lab filter
- Singer Lab (1) Apply Singer Lab filter
- Spruston Lab (1) Apply Spruston Lab filter
- Stern Lab (1) Apply Stern Lab filter
- Svoboda Lab (3) Apply Svoboda Lab filter
- Tjian Lab (1) Apply Tjian Lab filter
- Truman Lab (4) Apply Truman Lab filter
- Turaga Lab (1) Apply Turaga Lab filter
- Turner Lab (5) Apply Turner Lab filter
- Zuker Lab (1) Apply Zuker Lab filter
Associated Project Team
- CellMap (1) Apply CellMap filter
- Fly Functional Connectome (4) Apply Fly Functional Connectome filter
- Fly Olympiad (3) Apply Fly Olympiad filter
- FlyEM (12) Apply FlyEM filter
- FlyLight (20) Apply FlyLight filter
- GENIE (1) Apply GENIE filter
- Transcription Imaging (1) Apply Transcription Imaging filter
Publication Date
- 2026 (2) Apply 2026 filter
- 2025 (8) Apply 2025 filter
- 2024 (4) Apply 2024 filter
- 2023 (5) Apply 2023 filter
- 2022 (1) Apply 2022 filter
- 2021 (4) Apply 2021 filter
- 2020 (9) Apply 2020 filter
- 2019 (6) Apply 2019 filter
- 2018 (7) Apply 2018 filter
- 2017 (15) Apply 2017 filter
- 2016 (3) Apply 2016 filter
- 2015 (16) Apply 2015 filter
- 2014 (9) Apply 2014 filter
- 2013 (5) Apply 2013 filter
- 2012 (8) Apply 2012 filter
- 2011 (4) Apply 2011 filter
- 2010 (4) Apply 2010 filter
- 2009 (2) Apply 2009 filter
- 2008 (4) Apply 2008 filter
- 2007 (2) Apply 2007 filter
- 2006 (1) Apply 2006 filter
- 2002 (1) Apply 2002 filter
- 2000 (2) Apply 2000 filter
- 1999 (1) Apply 1999 filter
- 1997 (1) Apply 1997 filter
- 1995 (2) Apply 1995 filter
- 1994 (2) Apply 1994 filter
- 1993 (2) Apply 1993 filter
- 1992 (1) Apply 1992 filter
- 1991 (2) Apply 1991 filter
- 1990 (3) Apply 1990 filter
- 1989 (2) Apply 1989 filter
- 1987 (2) Apply 1987 filter
- 1986 (1) Apply 1986 filter
- 1985 (1) Apply 1985 filter
- 1984 (1) Apply 1984 filter
- 1983 (1) Apply 1983 filter
- 1982 (2) Apply 1982 filter
- 1981 (1) Apply 1981 filter
- 1979 (1) Apply 1979 filter
- 1973 (1) Apply 1973 filter
Type of Publication
149 Publications
Showing 61-70 of 149 resultsExogenous DNA sequences were introduced into the Drosophila germ line. A rosy transposon (ry1), constructed by inserting a chromosomal DNA fragment containing the wild-type rosy gene into a P transposable element, transformed germ line cells in 20 to 50 percent of the injected rosy mutant embryos. Transformants contained one or two copies of chromosomally integrated, intact ry1 that were stably inherited in subsequent generations. These transformed flies had wild-type eye color indicating that the visible genetic defect in the host strain could be fully and permanently corrected by the transferred gene. To demonstrate the generality of this approach, a DNA segment that does not confer a recognizable phenotype on recipients was also transferred into germ line chromosomes.
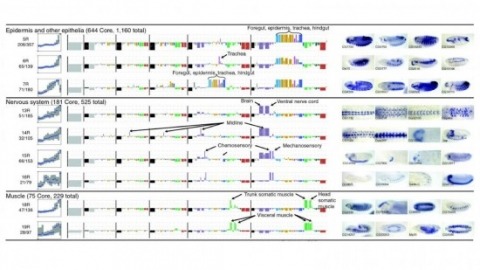
Cell and tissue specific gene expression is a defining feature of embryonic development in multi-cellular organisms. However, the range of gene expression patterns, the extent of the correlation of expression with function, and the classes of genes whose spatial expression are tightly regulated have been unclear due to the lack of an unbiased, genome-wide survey of gene expression patterns.
Although associative learning has been localized to specific brain areas in many animals, identifying the underlying synaptic processes in vivo has been difficult. Here, we provide the first demonstration of long-term synaptic plasticity at the output site of the Drosophila mushroom body. Pairing an odor with activation of specific dopamine neurons induces both learning and odor-specific synaptic depression. The plasticity induction strictly depends on the temporal order of the two stimuli, replicating the logical requirement for associative learning. Furthermore, we reveal that dopamine action is confined to and distinct across different anatomical compartments of the mushroom body lobes. Finally, we find that overlap between sparse representations of different odors defines both stimulus specificity of the plasticity and generalizability of associative memories across odors. Thus, the plasticity we find here not only manifests important features of associative learning but also provides general insights into how a sparse sensory code is read out.
We describe an engineered family of highly antigenic molecules based on GFP-like fluorescent proteins. These molecules contain numerous copies of peptide epitopes and simultaneously bind IgG antibodies at each location. These 'spaghetti monster' fluorescent proteins (smFPs) distributed well in neurons, notably into small dendrites, spines and axons. smFP immunolabeling localized weakly expressed proteins not well resolved with traditional epitope tags. By varying epitope and scaffold, we generated a diverse family of mutually orthogonal antigens. In cultured neurons and mouse and fly brains, smFP probes allowed robust, orthogonal multicolor visualization of proteins, cell populations and neuropil. smFP variants complement existing tracers and greatly increase the number of simultaneous imaging channels, and they performed well in advanced preparations such as array tomography, super-resolution fluorescence imaging and electron microscopy. In living cells, the probes improved single-molecule image tracking and increased yield for RNA-seq. These probes facilitate new experiments in connectomics, transcriptomics and protein localization.
Information processing relies on precise patterns of synapses between neurons. The cellular recognition mechanisms regulating this specificity are poorly understood. In the medulla of the Drosophila visual system, different neurons form synaptic connections in different layers. Here, we sought to identify candidate cell recognition molecules underlying this specificity. Using RNA sequencing (RNA-seq), we show that neurons with different synaptic specificities express unique combinations of mRNAs encoding hundreds of cell surface and secreted proteins. Using RNA-seq and protein tagging, we demonstrate that 21 paralogs of the Dpr family, a subclass of immunoglobulin (Ig)-domain containing proteins, are expressed in unique combinations in homologous neurons with different layer-specific synaptic connections. Dpr interacting proteins (DIPs), comprising nine paralogs of another subclass of Ig-containing proteins, are expressed in a complementary layer-specific fashion in a subset of synaptic partners. We propose that pairs of Dpr/DIP paralogs contribute to layer-specific patterns of synaptic connectivity.
The connectome provides large scale connectivity and morphology information for the majority of the central brain of . Using this data set, we provide a complete description of the olfactory system, covering all first, second and lateral horn-associated third-order neurons. We develop a generally applicable strategy to extract information flow and layered organisation from connectome graphs, mapping olfactory input to descending interneurons. This identifies a range of motifs including highly lateralised circuits in the antennal lobe and patterns of convergence downstream of the mushroom body and lateral horn. Leveraging a second data set we provide a first quantitative assessment of inter- versus intra-individual stereotypy. Comparing neurons across two brains (three hemispheres) reveals striking similarity in neuronal morphology across brains. Connectivity correlates with morphology and neurons of the same morphological type show similar connection variability within the same brain as across two brains.
Different types of Drosophila dopaminergic neurons (DANs) reinforce memories of unique valence and provide state-dependent motivational control [1]. Prior studies suggest that the compartment architecture of the mushroom body (MB) is the relevant resolution for distinct DAN functions [2, 3]. Here we used a recent electron microscope volume of the fly brain [4] to reconstruct the fine anatomy of individual DANs within three MB compartments. We find the 20 DANs of the γ5 compartment, at least some of which provide reward teaching signals, can be clustered into 5 anatomical subtypes that innervate different regions within γ5. Reconstructing 821 upstream neurons reveals input selectivity, supporting the functional relevance of DAN sub-classification. Only one PAM-γ5 DAN subtype (γ5fb) receives direct recurrent input from γ5β’2a mushroom body output neurons (MBONs) and behavioral experiments distinguish a role for these DANs in memory revaluation from those reinforcing sugar memory. Other DAN subtypes receive major, and potentially reinforcing, inputs from putative gustatory interneurons or lateral horn neurons, which can also relay indirect feedback from the MB. We similarly reconstructed the single aversively reinforcing PPL1-γ1pedc DAN. The γ1pedc DAN inputs are mostly different to those of γ5 DANs and are clustered onto distinct branches of its dendritic tree, presumably separating its established roles in aversive reinforcement and appetitive motivation [5, 6]. Additional tracing identified neurons that provide broad input to γ5, β’2a and γ1pedc DANs suggesting that distributed DAN populations can be coordinately regulated. These connectomic and behavioral analyses therefore reveal additional complexity of dopaminergic reinforcement circuits between and within MB compartments.
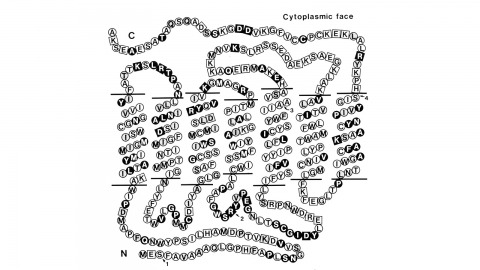
Using a novel method for detecting cross-homologous nucleic acid sequences we have isolated the gene coding for the major rhodopsin of Drosophila melanogaster and mapped it to chromosomal region 92B8-11. Comparison of cDNA and genomic DNA sequences indicates that the gene is divided into five exons. The amino acid sequence deduced from the nucleotide sequence is 373 residues long, and the polypeptide chain contains seven hydrophobic segments that appear to correspond to the seven transmembrane segments characteristic of other rhodopsins. Three regions of Drosophila rhodopsin are highly conserved with the corresponding domains of bovine rhodopsin, suggesting an important role for these polypeptide regions.
Janelia Farm, the new research campus of the Howard Hughes Medical Institute, is an ongoing experiment in the social engineering of research communities.
The development of the adult optic lobe (OL) of is directed by a wave of ingrowth of the photoreceptors over a two day period at the outset of metamorphosis which is accompanied by the appearance of the pupal-specific transcription factor Broad-Z3 (Br-Z3) and expression of early drivers in OL neurons. During this time, there are pulses of ecdysteroids that time the metamorphic events. At the outset, the transient appearance of juvenile hormone (JH) prevents precocious development of the OL caused by the ecdysteroid peak that initiates pupariation, but the artificial maintenance of JH after this time misdirects subsequent development. Axon ingrowth, Br-Z3 appearance and the expression of early drivers were unaffected, but aspects of later development such as the dendritic expansion of the lamina monopolar neurons and the expression of late drivers were suppressed. This effect of the exogenous JH mimic (JHM) pyriproxifen is lost by 24 hr after pupariation. Part of this effect of JHM is due to its suppression of the appearance of ecdysone receptor EcR-B1 that occurs after pupation and during early adult development.