Filter
Associated Lab
- Aguilera Castrejon Lab (4) Apply Aguilera Castrejon Lab filter
- Ahrens Lab (61) Apply Ahrens Lab filter
- Aso Lab (42) Apply Aso Lab filter
- Baker Lab (19) Apply Baker Lab filter
- Betzig Lab (104) Apply Betzig Lab filter
- Beyene Lab (10) Apply Beyene Lab filter
- Bock Lab (14) Apply Bock Lab filter
- Branson Lab (52) Apply Branson Lab filter
- Card Lab (37) Apply Card Lab filter
- Cardona Lab (45) Apply Cardona Lab filter
- Chklovskii Lab (10) Apply Chklovskii Lab filter
- Clapham Lab (15) Apply Clapham Lab filter
- Cui Lab (19) Apply Cui Lab filter
- Darshan Lab (8) Apply Darshan Lab filter
- Dennis Lab (2) Apply Dennis Lab filter
- Dickson Lab (32) Apply Dickson Lab filter
- Druckmann Lab (21) Apply Druckmann Lab filter
- Dudman Lab (44) Apply Dudman Lab filter
- Eddy/Rivas Lab (30) Apply Eddy/Rivas Lab filter
- Egnor Lab (4) Apply Egnor Lab filter
- Espinosa Medina Lab (19) Apply Espinosa Medina Lab filter
- Feliciano Lab (12) Apply Feliciano Lab filter
- Fetter Lab (31) Apply Fetter Lab filter
- FIB-SEM Technology (1) Apply FIB-SEM Technology filter
- Fitzgerald Lab (17) Apply Fitzgerald Lab filter
- Freeman Lab (15) Apply Freeman Lab filter
- Funke Lab (46) Apply Funke Lab filter
- Gonen Lab (59) Apply Gonen Lab filter
- Grigorieff Lab (34) Apply Grigorieff Lab filter
- Harris Lab (55) Apply Harris Lab filter
- Heberlein Lab (13) Apply Heberlein Lab filter
- Hermundstad Lab (26) Apply Hermundstad Lab filter
- Hess Lab (77) Apply Hess Lab filter
- Ilanges Lab (3) Apply Ilanges Lab filter
- Jayaraman Lab (44) Apply Jayaraman Lab filter
- Ji Lab (33) Apply Ji Lab filter
- Johnson Lab (1) Apply Johnson Lab filter
- Karpova Lab (13) Apply Karpova Lab filter
- Keleman Lab (8) Apply Keleman Lab filter
- Keller Lab (61) Apply Keller Lab filter
- Koay Lab (3) Apply Koay Lab filter
- Lavis Lab (146) Apply Lavis Lab filter
- Lee (Albert) Lab (29) Apply Lee (Albert) Lab filter
- Leonardo Lab (19) Apply Leonardo Lab filter
- Li Lab (8) Apply Li Lab filter
- Lippincott-Schwartz Lab (108) Apply Lippincott-Schwartz Lab filter
- Liu (Yin) Lab (3) Apply Liu (Yin) Lab filter
- Liu (Zhe) Lab (60) Apply Liu (Zhe) Lab filter
- Looger Lab (137) Apply Looger Lab filter
- Magee Lab (31) Apply Magee Lab filter
- Menon Lab (12) Apply Menon Lab filter
- Murphy Lab (6) Apply Murphy Lab filter
- O'Shea Lab (7) Apply O'Shea Lab filter
- Otopalik Lab (1) Apply Otopalik Lab filter
- Pachitariu Lab (40) Apply Pachitariu Lab filter
- Pastalkova Lab (6) Apply Pastalkova Lab filter
- Pavlopoulos Lab (7) Apply Pavlopoulos Lab filter
- Pedram Lab (4) Apply Pedram Lab filter
- Podgorski Lab (16) Apply Podgorski Lab filter
- Reiser Lab (49) Apply Reiser Lab filter
- Riddiford Lab (20) Apply Riddiford Lab filter
- Romani Lab (39) Apply Romani Lab filter
- Rubin Lab (111) Apply Rubin Lab filter
- Saalfeld Lab (47) Apply Saalfeld Lab filter
- Satou Lab (3) Apply Satou Lab filter
- Scheffer Lab (38) Apply Scheffer Lab filter
- Schreiter Lab (53) Apply Schreiter Lab filter
- Schulze Lab (1) Apply Schulze Lab filter
- Sgro Lab (3) Apply Sgro Lab filter
- Shroff Lab (31) Apply Shroff Lab filter
- Simpson Lab (18) Apply Simpson Lab filter
- Singer Lab (37) Apply Singer Lab filter
- Spruston Lab (62) Apply Spruston Lab filter
- Stern Lab (77) Apply Stern Lab filter
- Sternson Lab (47) Apply Sternson Lab filter
- Stringer Lab (38) Apply Stringer Lab filter
- Svoboda Lab (132) Apply Svoboda Lab filter
- Tebo Lab (11) Apply Tebo Lab filter
- Tervo Lab (9) Apply Tervo Lab filter
- Tillberg Lab (19) Apply Tillberg Lab filter
- Tjian Lab (17) Apply Tjian Lab filter
- Truman Lab (58) Apply Truman Lab filter
- Turaga Lab (41) Apply Turaga Lab filter
- Turner Lab (27) Apply Turner Lab filter
- Vale Lab (8) Apply Vale Lab filter
- Voigts Lab (4) Apply Voigts Lab filter
- Wang (Meng) Lab (28) Apply Wang (Meng) Lab filter
- Wang (Shaohe) Lab (6) Apply Wang (Shaohe) Lab filter
- Wong-Campos Lab (1) Apply Wong-Campos Lab filter
- Wu Lab (8) Apply Wu Lab filter
- Zlatic Lab (26) Apply Zlatic Lab filter
- Zuker Lab (5) Apply Zuker Lab filter
Associated Project Team
- CellMap (13) Apply CellMap filter
- COSEM (3) Apply COSEM filter
- FIB-SEM Technology (5) Apply FIB-SEM Technology filter
- Fly Descending Interneuron (12) Apply Fly Descending Interneuron filter
- Fly Functional Connectome (14) Apply Fly Functional Connectome filter
- Fly Olympiad (5) Apply Fly Olympiad filter
- FlyEM (56) Apply FlyEM filter
- FlyLight (50) Apply FlyLight filter
- GENIE (47) Apply GENIE filter
- Integrative Imaging (9) Apply Integrative Imaging filter
- Larval Olympiad (2) Apply Larval Olympiad filter
- MouseLight (18) Apply MouseLight filter
- NeuroSeq (1) Apply NeuroSeq filter
- ThalamoSeq (1) Apply ThalamoSeq filter
- Tool Translation Team (T3) (29) Apply Tool Translation Team (T3) filter
- Transcription Imaging (45) Apply Transcription Imaging filter
Associated Support Team
- Project Pipeline Support (5) Apply Project Pipeline Support filter
- Anatomy and Histology (18) Apply Anatomy and Histology filter
- Cryo-Electron Microscopy (44) Apply Cryo-Electron Microscopy filter
- Electron Microscopy (18) Apply Electron Microscopy filter
- Gene Targeting and Transgenics (11) Apply Gene Targeting and Transgenics filter
- High Performance Computing (7) Apply High Performance Computing filter
- Integrative Imaging (23) Apply Integrative Imaging filter
- Invertebrate Shared Resource (40) Apply Invertebrate Shared Resource filter
- Janelia Experimental Technology (37) Apply Janelia Experimental Technology filter
- Management Team (1) Apply Management Team filter
- Mass Spectrometry (1) Apply Mass Spectrometry filter
- Molecular Genomics (15) Apply Molecular Genomics filter
- Project Technical Resources (54) Apply Project Technical Resources filter
- Quantitative Genomics (20) Apply Quantitative Genomics filter
- Scientific Computing (103) Apply Scientific Computing filter
- Stem Cell & Primary Culture (14) Apply Stem Cell & Primary Culture filter
- Viral Tools (14) Apply Viral Tools filter
- Vivarium (7) Apply Vivarium filter
Publication Date
- 2026 (65) Apply 2026 filter
- 2025 (222) Apply 2025 filter
- 2024 (210) Apply 2024 filter
- 2023 (157) Apply 2023 filter
- 2022 (166) Apply 2022 filter
- 2021 (175) Apply 2021 filter
- 2020 (177) Apply 2020 filter
- 2019 (177) Apply 2019 filter
- 2018 (206) Apply 2018 filter
- 2017 (186) Apply 2017 filter
- 2016 (191) Apply 2016 filter
- 2015 (195) Apply 2015 filter
- 2014 (190) Apply 2014 filter
- 2013 (136) Apply 2013 filter
- 2012 (112) Apply 2012 filter
- 2011 (98) Apply 2011 filter
- 2010 (61) Apply 2010 filter
- 2009 (56) Apply 2009 filter
- 2008 (40) Apply 2008 filter
- 2007 (21) Apply 2007 filter
- 2006 (3) Apply 2006 filter
2844 Janelia Publications
Showing 81-90 of 2844 resultsExtracellular expression of heat shock protein 90 (eHsp90) by tumor cells is correlated with malignancy. Development of small molecule probes that can detect eHsp90 in vivo may therefore have utility in the early detection of malignancy. We synthesized a cell impermeable far-red fluorophore-tagged Hsp90 inhibitor to target eHsp90 in vivo. High resolution confocal and lattice light sheet microscopy show that probe-bound eHsp90 accumulates in punctate structures on the plasma membrane of breast tumor cells and is actively internalized. The extent of internalization correlates with tumor cell aggressiveness, and this process can be induced in benign cells by over-expressing p110HER2. Whole body cryoslicing, imaging and histology of flank and spontaneous tumor-bearing mice strongly suggests that eHsp90 expression and internalization is a phenomenon unique to tumor cells in vivo and may provide an 'Achilles heel' for the early diagnosis of metastatic disease and targeted drug delivery.
Contact sites between lipid droplets and other organelles are essential for cellular lipid and energy homeostasis upon metabolic demands. Detection of these contact sites at the nanometer scale over time in living cells is challenging. We developed a tool kit for detecting contact sites based on fluorogen-activated bimolecular complementation at CONtact sites, FABCON, using a reversible, low-affinity split fluorescent protein, splitFAST. FABCON labels contact sites with minimal perturbation to organelle interaction. Via FABCON, we quantitatively demonstrated that endoplasmic reticulum (ER)- and mitochondria (mito)-lipid droplet contact sites are dynamic foci in distinct metabolic conditions, such as during lipid droplet biogenesis and consumption. An automated analysis pipeline further classified individual contact sites into distinct subgroups based on size, likely reflecting differential regulation and function. Moreover, FABCON is generalizable to visualize a repertoire of organelle contact sites including ER-mito. Altogether, FABCON reveals insights into the dynamic regulation of lipid droplet-organelle contact sites and generates new hypotheses for further mechanistical interrogation during metabolic regulation.
Although different animal species often exhibit extensive variation in many behaviors, typically scientists examine one or a small number of behaviors in any single study. Here, we propose a new framework to simultaneously study the evolution of many behaviors. We measured the behavioral repertoire of individuals from six species of fruit flies using unsupervised techniques and identified all stereotyped movements exhibited by each species. We then fit a Generalized Linear Mixed Model to estimate the intra- and inter-species behavioral covariances, and, by using the known phylogenetic relationships among species, we estimated the (unobserved) behaviors exhibited by ancestral species. We found that much of intra-specific behavioral variation has a similar covariance structure to previously described long-time scale variation in an individual's behavior, suggesting that much of the measured variation between individuals of a single species in our assay reflects differences in the status of neural networks, rather than genetic or developmental differences between individuals. We then propose a method to identify groups of behaviors that appear to have evolved in a correlated manner, illustrating how sets of behaviors, rather than individual behaviors, likely evolved. Our approach provides a new framework for identifying co-evolving behaviors and may provide new opportunities to study the mechanistic basis of behavioral evolution.
Regions of genomic DNA called enhancers encode binding sites for transcription factor proteins. Binding of activators and repressors increase and reduce transcription, respectively, but it is not understood how combinations of activators and repressors generate precise patterns of transcription during development. Here, we explore this problem using a fully synthetic transcriptional platform in Drosophila consisting of engineered transcription factor gradients and artificial enhancers. We found that binding sites for a transcription factor that makes DNA accessible are required together with binding sites for transcriptional activators to produce a functional enhancer. Only in this context can changes in the number of activator binding sites mediate quantitative control of transcription. Using an engineered transcriptional repressor gradient, we demonstrate that overlapping repressor and activator binding sites provide more robust repression and sharper expression boundaries than non-overlapping sites. This may explain why this common motif is observed in many developmental enhancers.
Topographic maps, the systematic spatial ordering of neurons by response tuning, are common across species. In Drosophila, the lobula columnar (LC) neuron types project from the optic lobe to the central brain, where each forms a glomerulus in a distinct position. However, the advantages of this glomerular arrangement are unclear. Here, we examine the functional and spatial relationships of 10 glomeruli using single-neuron calcium imaging. We discover novel detectors for objects smaller than the lens resolution (LC18) and for complex line motion (LC25). We find that glomeruli are spatially clustered by selectivity for looming versus drifting object motion and ordered by size tuning to form a topographic visual feature map. Furthermore, connectome analysis shows that downstream neurons integrate from sparse subsets of possible glomeruli combinations, which are biased for glomeruli encoding similar features. LC neurons are thus an explicit example of distinct feature detectors topographically organized to facilitate downstream circuit integration.
We report the larval CNS expression patterns for 6,650 GAL4 lines based on cis-regulatory regions (CRMs) from the Drosophila genome. Adult CNS expression patterns were previously reported for this collection, thereby providing a unique resource for determining the origins of adult cells. An illustrative example reveals the origin of the astrocyte-like glia of the ventral CNS. Besides larval neurons and glia, the larval CNS contains scattered lineages of immature, adult-specific neurons. Comparison of lineage expression within this large collection of CRMs provides insight into the codes used for designating neuronal types. The CRMs encode both dense and sparse patterns of lineage expression. There is little correlation between brain and thoracic lineages in patterns of sparse expression, but expression in the two regions is highly correlated in the dense mode. The optic lobes, by comparison, appear to use a different set of genetic instructions in their development.
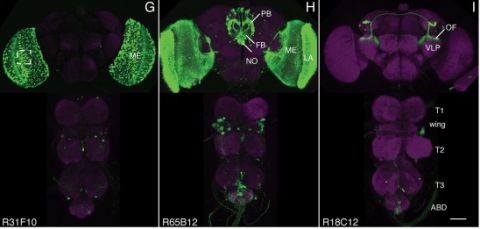
We established a collection of 7,000 transgenic lines of Drosophila melanogaster. Expression of GAL4 in each line is controlled by a different, defined fragment of genomic DNA that serves as a transcriptional enhancer. We used confocal microscopy of dissected nervous systems to determine the expression patterns driven by each fragment in the adult brain and ventral nerve cord. We present image data on 6,650 lines. Using both manual and machine-assisted annotation, we describe the expression patterns in the most useful lines. We illustrate the utility of these data for identifying novel neuronal cell types, revealing brain asymmetry, and describing the nature and extent of neuronal shape stereotypy. The GAL4 lines allow expression of exogenous genes in distinct, small subsets of the adult nervous system. The set of DNA fragments, each driving a documented expression pattern, will facilitate the generation of additional constructs for manipulating neuronal function. synapse was substantially elevated, at the endocytic zone there was no enhanced polymerization activity. We conclude that actin subserves spatially diverse, independently regulated processes throughout spines. Perisynaptic actin forms a uniquely dynamic structure well suited for direct, active regulation of the synapse.
For the overall strategy and methods used to produce the GAL4 lines:
Pfeiffer, B.D., Jenett, A., Hammonds, A.S., Ngo, T.T., Misra, S., Murphy, C., Scully, A., Carlson, J.W., Wan, K.H., Laverty, T.R., Mungall, C., Svirskas, R., Kadonaga, J.T., Doe, C.Q., Eisen, M.B., Celniker, S.E., Rubin, G.M. (2008). Tools for neuroanatomy and neurogenetics in Drosophila. Proc. Natl. Acad. Sci. USA 105, 9715-9720. http://www.pnas.org/content/105/28/9715.full.pdf+html synapse was substantially elevated, at the endocytic zone there was no enhanced polymerization activity. We conclude that actin subserves spatially diverse, independently regulated processes throughout spines. Perisynaptic actin forms a uniquely dynamic structure well suited for direct, active regulation of the synapse.
For data on expression in the embryo:
Manning, L., Purice, M.D., Roberts, J., Pollard, J.L., Bennett, A.L., Kroll, J.R., Dyukareva, A.V., Doan, P.N., Lupton, J.R., Strader, M.E., Tanner, S., Bauer, D., Wilbur, A., Tran, K.D., Laverty, T.R., Pearson, J.C., Crews, S.T., Rubin, G.M., and Doe, C.Q. (2012) Annotated embryonic CNS expression patterns of 5000 GMR GAL4 lines: a resource for manipulating gene expression and analyzing cis-regulatory motifs. Cell Reports (2012) Doi: 10.1016/j.celrep.2012.09.009 http://www.cell.com/cell-reports/fulltext/S2211-1247(12)00290-2 synapse was substantially elevated, at the endocytic zone there was no enhanced polymerization activity. We conclude that actin subserves spatially diverse, independently regulated processes throughout spines. Perisynaptic actin forms a uniquely dynamic structure well suited for direct, active regulation of the synapse.
For data on expression in imaginal discs:
Jory, A., Estella, C., Giorgianni, M.W., Slattery, M., Laverty, T.R., Rubin, G.M., and Mann, R.S. (2012) A survey of 6300 genomic fragments for cis-regulatory activity in the imaginal discs of Drosophila melanogaster. Cell Reports (2012) Doi: 10.1016/j.celrep.2012.09.010 http://www.cell.com/cell-reports/fulltext/S2211-1247(12)00291-4 synapse was substantially elevated, at the endocytic zone there was no enhanced polymerization activity. We conclude that actin subserves spatially diverse, independently regulated processes throughout spines. Perisynaptic actin forms a uniquely dynamic structure well suited for direct, active regulation of the synapse.
For data on expression in the larval nervous system:
Li, H.-H., Kroll, J.R., Lennox, S., Ogundeyi, O., Jeter, J., Depasquale, G., and Truman, J.W. (2013) A GAL4 driver resource for developmental and behavioral studies on the larval CNS of Drosophila. Cell Reports (submitted).
Fruit flies recognize hundreds of ecologically relevant odors and respond appropriately to them. The complexity, redundancy and interconnectedness of the olfactory machinery complicate efforts to pinpoint the functional contributions of any component neuron or receptor to behavior. Some contributions can only be elucidated in flies that carry multiple mutations and transgenes, but the production of such flies is currently labor-intensive and time-consuming. Here, we describe a set of transgenic flies that express the GAL80 in specific olfactory sensory neurons (). The GAL80s effectively and specifically subtract the activities of GAL4-driven transgenes that impart anatomical and physiological phenotypes. can allow researchers to efficiently activate only one or a few types of functional neurons in an otherwise nonfunctional olfactory background. Such experiments will improve our understanding of the mechanistic connections between odorant inputs and behavioral outputs at the resolution of only a few functional neurons.
We engineered electrochromic fluorescence resonance energy transfer (eFRET) genetically encoded voltage indicators (GEVIs) with “positive-going” fluorescence response to membrane depolarization through rational manipulation of the native proton transport pathway in microbial rhodopsins. We transformed the state-of-the-art eFRET GEVI Voltron into Positron, with kinetics and sensitivity equivalent to Voltron but flipped fluorescence signal polarity. We further applied this general approach to GEVIs containing different voltage sensitive rhodopsin domains and various fluorescent dye and fluorescent protein reporters.
Pushing the frontier of fluorescence microscopy requires the design of enhanced fluorophores with finely tuned properties. We recently discovered that incorporation of four-membered azetidine rings into classic fluorophore structures elicits substantial increases in brightness and photostability, resulting in the Janelia Fluor (JF) series of dyes. We refined and extended this strategy, finding that incorporation of 3-substituted azetidine groups allows rational tuning of the spectral and chemical properties of rhodamine dyes with unprecedented precision. This strategy allowed us to establish principles for fine-tuning the properties of fluorophores and to develop a palette of new fluorescent and fluorogenic labels with excitation ranging from blue to the far-red. Our results demonstrate the versatility of these new dyes in cells, tissues and animals.